Welcome to the Biomedical Research Computing Facility (BRCF) Users wiki! Formerly known as the Research Computing Task Force (RCTF), the BRCF now has an official organizational home in the Center for Biomedical Research Support (CBRS).
Our mission: Biomedical Research Computing Facility (BRCF) "PODs" (small compute clusters) provide a standard hardware, software and storage architecture, suitable for local, interactive biomedical research computing, that can be efficiently managed.
Upcoming Maintenance
POD maintenance schedule for April 2024
- BRCF research PODs: 8am - 6pm Tuesday April 16
- No EDU pod maintenance
Python version updates
After the January 2024 maintenance JupyterHub Python version is now 3.9 on all compute servers. Python 3.9 is also available on the command line by calling it explicitly: i.e., python3.9; the default command line python3 version is still 3.8.
Many Python packages were also updated, in both the Python 3.8 and 3.9 command line environments as well as in the JupyterHub Python 3.8 environment. Please contact us at rctf-support@utexas.edu if you experience Python-related issues.
See this Wiki page for more information about Python on BRCF pods: About Python and JupyterHub server
Quick links
- Architecture Overview and Goals
- Obtaining a BRCF Account (https://rctf-account-request.icmb.utexas.edu)
- Accessible only from the UT campus network or with the UT VPN service active.
- POD Resources and Access
- Support and Maintenance
- POD Software Information
- The Educational POD
Architecture Overview
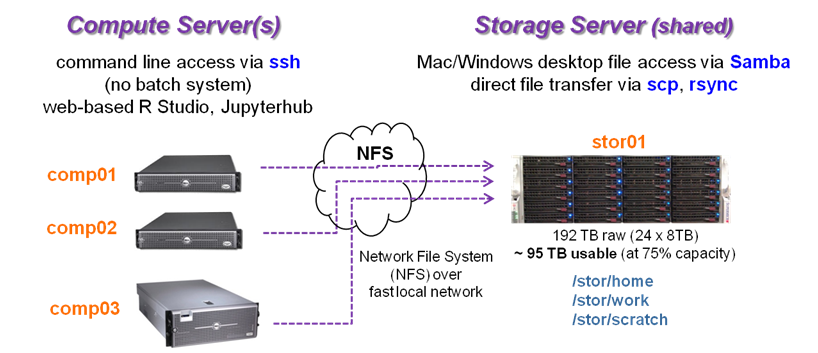
BRCF provides local, centralized storage and compute systems called PODs. A POD consists of one or more compute servers along with a shared storage server. Files on POD storage can be accessed from any server within that POD.
A graphic illustration of the BRCF POD Compute/Storage model is shown below:
Features of this architecture include:
- A large set of Bioinformatics software available on all compute servers
- interactive (non-batch) environment supports exploratory analyses and long-running jobs
- web-based R Studio and JupyterHub servers implemented on all compute servers
- Storage managed by the high-performance ZFS file system, with
- built-in data integrity, superior to standard Unix file systems
- large, contiguous address space
- automatic file compression
- RAID-like redundancy capabilities
- works well with inexpensive, commodity disks
- Appropriate organization and management of research data
- weekly backup of Home and Work to spinning disk at the UT Data Center (UDC)
- periodic archiving of backup data to TACC's ranch tape archive system (~yearly)
- Centralized deployment, administration and monitoring
- OS configuration and 3rd part software installed via the Puppet deployment tool
- global BRCF user and group IDs, deployable to any POD
- self-service Account Management web interface
- OS monitoring via Nagios tool
- hardware-level monitoring via Out-Of-Band Management (OOBM) interfaces
- IMPI, iDRAC
BRCF Architecture goals
The Biomedical Research Computing Facility (BRCF) is a core facility in UT's Center for Biomedical Research Support (CBRS), under the Office of the Vice President of Research. Our small compute clusters, known as "PODs", provide a standard hardware, software storage architecture, suitable for local research computing, that can be efficiently managed.
The BRCF grew out of the earlier Research Computing Task Force (RCTF) working group of IT-knowledgeable UT staff and students from CSSB, GSAF and CCBB. Today OurTeam consists of staff from Molecular Biosciences, CBRS, and the College of Natural Sciences Office of Information Technology (CNS-OIT).
Broadly, our goals are to supplement TACC's offerings by providing extensive local storage, including backups and archiving, along with easy-access non-batch local compute.
Before the BRCF initiative, labs had their own legacy computational equipment and storage, as well as a hodgepodge of backup solutions (a common solution being "none"). This diversity combined with dwindling systems administration resources led to an untenable situation.
The Texas Advanced Computing Center (TACC) provides excellent computation resources for performing large-scale computations in parallel. However its batch system orientation is not well suited for running smaller, one-off computations, for developing scripts and pipelines, or for executing very long-running (> 2 days) computations. While TACC offers a no-cost tape archive facility (ranch), its persistent storage offerings (corral, global work file system) can be cumbersome to use for collaboration.
The BRCF POD architecture has been designed to address these issues and needs.
- Provide adequate local storage (spinning disk) in a large, non-partitioned address space.
- Implement some common file system structures to assist data organization and automation
- Provide flexible local compute capability with both common and lab-specific bioinformatics tools installed.
- Augment TACC offerings with non-batch computing environment
- Be robust and "highly available" (but 24x7 uptime not required)
- Provide automated backups to spinning disk and periodic data archiving to TACC's ranch tape system.
- Target the "sweet spot" of cost-versus-function commodity hardware offerings
- Aim for rolling hardware upgrades as technology evolves
- Provide centralized management of POD equipment
- Automate software and configuration deployment and system monitoring
- Make it easy to deploy new equipment
Recent Changes
R version updates
After the August 15, 2023 maintenance the default R version is R 4.3.1 both for RStudio Server on all compute servers, and on the command line. See About R and R Studio Server for more information.
OS upgrade changes
In March 2023, the BRCF completed upgrading the Operating System (OS) software on all our POD servers. See this page for important information about how this change may affect you: Winter 22-23 OS Upgrades.
Change to SSH key access
After the Tuesday June 28, 2022 maintenance, remote SSH access to POD compute servers no longer uses non-standard port 222. Instead, uses standard port 22, so no port (-p) option is required in the ssh command.
Recently Updated
Navigate space
Recent space activity