| Tip |
|---|
|
Use our summer school reservation (CoreNGS-Thu) when submitting batch jobs to get higher priority on the ls6 normal queue today: sbatch --reservation=CoreNGS-Thu <batch_file>.slurm
idev -m 180 -N 1 -A OTH21164 -r CoreNGS-Thu
|
Overview
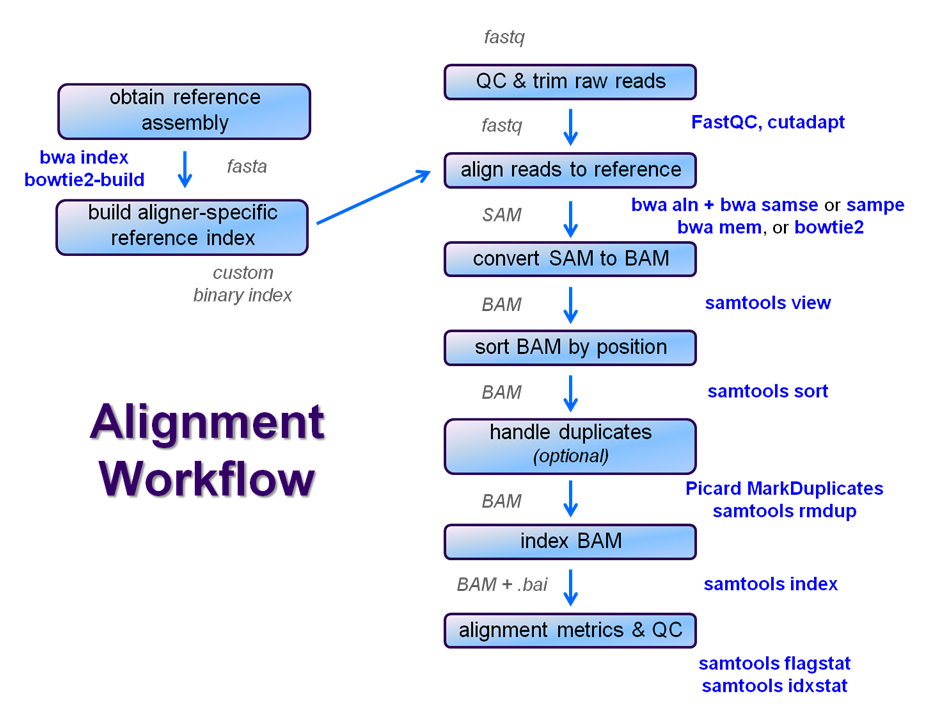 Image Removed
Image Removed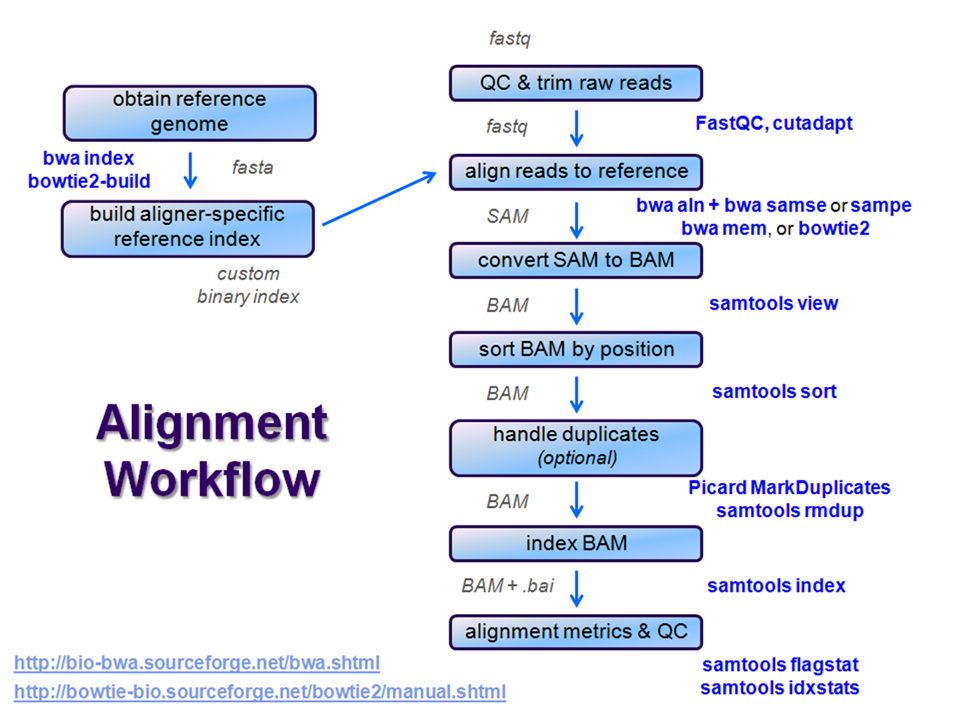 Image Added
Image Added
After raw sequence files are generated (in FASTQ format), quality-checked, and pre-processed in some way, the next step in many NGS pipelines is mapping to a reference genome.
...
Even though many mapping tools exist, a few individual programs have a dominant "market share" of the NGS world. In this section, we will primarily focus on two of the most versatile general-purpose ones: BWA and Bowtie2 (the latter being part of the Tuxedo suite which includes the transcriptome-aware RNA-seq aligner Tophat2 as well as other downstream quantifiaction quantification tools).
Stage the alignment data
First connect to stampede2ls6.tacc.utexas.edu and start an idev session. This should be second nature by now 
| Code Block |
|---|
| language | bash |
|---|
| title | Start an idev session |
|---|
|
idev -pm normal180 -mN 1801 -A UT-2015-05-18OTH21164 -Nr 1 -n 68 CoreNGS-Thu |
Then stage the sample datasets and references we will use.
| Code Block |
|---|
| language | bash |
|---|
| title | Get the alignment exercises files |
|---|
|
mkdir -p $SCRATCH/core_ngs/references/fasta# Copy the FASTA files for building references
mkdir -p $SCRATCH/core_ngs/alignmentreferences/fastqfasta
cp $CORENGS/references/fasta/*.fa $SCRATCH/core_ngs/references/fasta/
cp
# Copy the FASTQ files that will be used for alignment
mkdir -p $SCRATCH/core_ngs/alignment/fastq
cp $CORENGS/alignment/*fastq.gz $SCRATCH/core_ngs/alignment/fastq/
cd $SCRATCH/core_ngs/alignment/fastq |
...
| File Name | Description | Sample |
|---|
| Sample_Yeast_L005_R1.cat.fastq.gz | Paired-end Illumina, First of pair, FASTQ | Yeast ChIP-seq |
| Sample_Yeast_L005_R2.cat.fastq.gz | Paired-end Illumina, Second of pair, FASTQ | Yeast ChIP-seq |
| human_rnaseq.fastq.gz | Paired-end Illumina, First of pair only, FASTQ | Human RNA-seq |
| human_mirnaseq.fastq.gz | Single-end Illumina, FASTQ | Human microRNA-seq |
| cholera_rnaseq.fastq.gz | Single-end Illumina, FASTQ | V. cholerae RNA-seq |
Reference Genomes
...
Here are the four reference genomes we will be using today, with some information about them. These are not necessarily the most recent versions of these references (e.g. the newest human reference genome is hg38 and the most a recent miRBase annotation is version is v21. (See here for information about many more genomes.)
...
Searching genomes is computationally hard work and takes a long time if done on linear genomic sequence. So aligners require that references first be indexed to accelerate lookup. The aligners we are using each require a different index, but use the same method (the Burrows-Wheeler Transform) to get the job done.
Building a reference index involves taking a FASTA file as input, with each contig (contiguous string of bases, e.g. a chromosome) as a separate FASTA entry, and producing an aligner-specific set of files as output. Those output index files are then used to perform the sequence alignment, and alignments are reported using coordinates referencing names and offset positions based on the original FASTA file contig entries.
...
| Code Block |
|---|
| language | bash |
|---|
| title | BWA hg19 index location |
|---|
|
/work2work/projects/BioITeam/ref_genome/bwa/bwtsw/hg19 |
...
| Tip |
|---|
The BioITeam maintains a set of reference indexes for many common organisms and aligners. They can be found in aligner-specific sub-directories of the /work2work/projects/BioITeam/ref_genome area. E.g.: | Code Block |
|---|
| /work2work/projects/BioITeam/ref_genome/
bowtie2/
bwa/
hisat2/
kallisto/
star/
tophat/ |
|
...
We've discovered a pattern (also known as a regular expression) to use in searching, and the command line tool that does regular expression matching is grep (general regular expression parser). (Read more about grep here: Advanced commands: grep.and regular expressions)
Regular expressions are so powerful that nearly every modern computer language includes a "regex" module of some sort. There are many online tutorials for regular expressions, and several slightly different "flavors" of them. But the most common is the Perl style (http://perldoc.perl.org/perlretut.html), which was one of the fist and still the most powerful (there's a reason Perl was used extensively when assembling the human genome). We're only going to use the most simple of regular expressions here, but learning more about them will pay handsome dividends for you in the future.
...
| Code Block |
|---|
| language | bash |
|---|
| title | grep to match contig names in a FASTA file |
|---|
|
# Stage the FASTA files
cds
mkdir -p core_ngs/references/fasta
cd core_ngs/references/fasta
cp $CORENGS/references/fasta/*.fa .
cd $SCRATCH/core_ngs/references/fasta
grep -P '^>' sacCer3.fa | more |
...
Now down to the nuts and bolts of the pattern: '^>^>'
First, the single quotes around the pattern – this tells the bash shell to pass the exact string contents to grep.
As part of its friendly we have seen, during command line parsing and evaluation , the shell will often look for special characters metacharacters on the command line that mean something to it (for example, the the $ in front of an environment variable name, like in $SCRATCH). Well, regular expressions treat the $ specially too – but in a completely different way! Those Those single quotes tell tell the shell "don't look inside here for special characters – treat this as a literal string and pass it to the program". The shell will obey, will strip the single quotes off the string, and will pass the actual pattern, ^> ^>, to the grep program. (Note that the shell does look inside double quotes ( " ) for certain special signals, such as looking for environment variable names to evaluate. Read more about program. (Read more about Quoting in the shell.)
So what does ^>^> mean to grep? We know that contig name lines always start with a > character, so > is a literal for grep to use in its pattern match.
We might be able to get away with just using this literal alone as our regex, specifying '>' as as the command line pattern argument. But for for grep, the more specific the pattern, the better. So we constrain where the > can appear on the line. The special carat ( ^ ) metacharacter represents "beginning of line". So ^>^> means "beginning of a line followed by a > character".Exercise: How many contigs are there in the sacCer3 reference?
| Expand |
|---|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Get the alignment exercises files |
|---|
| # Copy the FASTA files for building references
mkdir -p $SCRATCH/core_ngs/references/fasta
cp $CORENGS/references/fasta/*.fa $SCRATCH/core_ngs/references/fasta/ |
|
Exercise: How many contigs are there in the sacCer3 reference?
| Expand |
|---|
|
| Code Block |
|---|
| cd $SCRATCH/core_ngs/references/fasta
grep -P '^>' sacCer3.fa | wc -l |
Or use grep's -c option that says "just count the line matches" | Code Block |
|---|
| grep -P -c '^>' sacCer3.fa |
|
...
| alignment type | aligner options | pro's | con's |
|---|
| global with bwa | SEsingle end reads: PEpaired end reads: - bwa aln <R1>
- bwa aln <R2>
- bwa sampe
| - simple to use (take default options)
- good for basic global alignment
| |
| global with bowtie2bowtie2 --global | bowtie2 | - extremely configurable
- can be used for RNAseq alignment (after adapter trimming) because of its many options
| |
| local with bwa | bwa mem | - simple to use (take default options)
- very fast
- no adapter trimming needed
- good for simple RNAseq analysis
- the secondary alignments it reports provide splice junction information
| - always produces alignments with secondary reads
- must be filtered if not desired
|
| local with bowtie2 | bowtie2 --local | - extremely configurable
- no adapter trimming needed
- good for small RNA alignment because of its many options
| |
...
- Trim the FASTQ sequences down to 50 with fastx_clipper
- this removes most of any 5' adapter contamination without the fuss of specific adapter trimming w/cutadapt
- Prepare the sacCer3 reference index for bwa using bwa index
- this is done once, and re-used for later alignments
- Perform a global bwa alignment on the R1 reads (bwa aln) producing a BWA-specific binary .sai intermediate file
- Perform a global bwa alignment on the R2 reads (bwa aln) producing a BWA-specific binary .sai intermediate file
- Perform pairing of the separately aligned reads and report the alignments in SAM format using bwa sampe
- Convert the SAM file to a BAM file (samtools view)
- Sort the BAM file by genomic location (samtools sort)
- Index the BAM file (samtools index)
- Gather simple alignment statistics (samtools flagstat and samtools idxstatidxstats)
We're going to skip the trimming step for now and see how it goes. We'll perform steps 2 - 5 now and leave samtools for a later exercise since steps 6 - 10 are common to nearly all post-alignment workflows.
...
Like other tools you've worked with so far, you first need to load bwa. Do that now, and then enter bwa with no arguments to view the top-level help page (many NGS tools will provide some help when called with no arguments). Note that bwa is available both from the standard TACC module system and as as a BioContainers. module.
...
Make sure you're in
...
an idev session
| Code Block |
|---|
| language | bash |
|---|
| title | Start an idev session |
|---|
|
idev - pm normal180 - mN 1201 -A OTH21164 UT- 2015-05-18r CoreNGS-Thu
# or
idev -m 90 -N 1 - n 68 A OTH21164 -p development |
| Code Block |
|---|
|
module load biocontainers # takes a while
module load bwa
bwa |
...
| Expand |
|---|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Get the alignment exercises files |
|---|
| mkdir -p $SCRATCH/core_ngs/alignment/fastq
mkdir -p $SCRATCH/core_ngs/references/fasta
cp $CORENGS/alignment/*fastq.gz $SCRATCH/core_ngs/alignment/fastq/
cp $CORENGS/references/fasta/*.fa $SCRATCH/core_ngs/references/fasta/ |
|
...
| Code Block |
|---|
| language | bash |
|---|
| title | Prepare BWA reference directory for sacCer3 |
|---|
|
mkdir -p $SCRATCH/core_ngs/references/bwa/sacCer3
cd $SCRATCH/core_ngs/references/bwa/sacCer3
ln -ssf ../../fasta/sacCer3.fa
ls -l |
...
| Expand |
|---|
|
| Code Block |
|---|
| # Copy the pre-built FASTA files for building references
mkdir -p $SCRATCH/core_ngs/references
cp $CORENGS/references/fasta/*.fa $SCRATCH/core_ngs/references/fasta/
# Get the FASTQ to align Copy a pre-built bwa index for sacCer3
mkdir -p $SCRATCH/core_ngs/references/alignmentbwa/fastqsacCer3
cp $CORENGS/references/bwa/alignmentsacCer3/*fastq.gz* $SCRATCH/core_ngs/references/alignmentbwa/fastqsacCer3/
# Get the FASTQ to align
mkdir -p $SCRATCH/core_ngs/alignment/fastq
cp $CORENGS/alignment/*fastq.gz $SCRATCH/core_ngs/alignment/fastq/
|
|
| Code Block |
|---|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Prepare to align yeast data |
|---|
|
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
ln -s -fsf ../fastq
ln -s -fsf ../../references/bwa/sacCer3 |
...
Required arguments are a <prefix> of the bwa index files, and the input FASTQ file. There are lots of options, but here is a summary of the most important ones.
...
| Code Block |
|---|
| language | bash |
|---|
| title | bwa aln commands for yeast R1 and R2 |
|---|
|
# If not already loaded:
module load biocontainers
module load bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
bwa aln sacCer3/sacCer3.fa fastq/Sample_Yeast_L005_R1.cat.fastq.gz > yeast_pe_R1.sai
bwa aln sacCer3/sacCer3.fa fastq/Sample_Yeast_L005_R2.cat.fastq.gz > yeast_pe_R2.sai |
When all is done you should have two .sai files: yeast_pe_R1.sai and yeast_pe_R2.sai.
| Tip |
|---|
| title | Make sure your output files are not empty |
|---|
|
Double check that output was written by doing ls -lh and making sure the file sizes listed are not 0. |
...
| Expand |
|---|
|
The last few lines of bwa's execution output should look something like this: | Code Block |
|---|
| [bwa_aln] 17bp reads: max_diff = 2
[bwa_aln] 38bp reads: max_diff = 3
[bwa_aln] 64bp reads: max_diff = 4
[bwa_aln] 93bp reads: max_diff = 5
[bwa_aln] 124bp reads: max_diff = 6
[bwa_aln] 157bp reads: max_diff = 7
[bwa_aln] 190bp reads: max_diff = 8
[bwa_aln] 225bp reads: max_diff = 9
[bwa_aln_core] calculate SA coordinate... 50.76 sec
[bwa_aln_core] write to the disk... 0.07 sec
[bwa_aln_core] 262144 sequences have been processed.
[bwa_aln_core] calculate SA coordinate... 50.35 sec
[bwa_aln_core] write to the disk... 0.07 sec
[bwa_aln_core] 524288 sequences have been processed.
[bwa_aln_core] calculate SA coordinate... 13.64 sec
[bwa_aln_core] write to the disk... 0.01 sec
[bwa_aln_core] 592180 sequences have been processed.
[main] Version: 0.7.17-r1188
[main] CMD: /usr/local/bin/bwa aln sacCer3/sacCer3.fa fastq/Sample_Yeast_L005_R1.cat.fastq.gz
[main] Real time: 12278.936185 sec; CPU: 12377.597598 sec
|
So the R2 alignment took ~123 ~78 seconds (~2 ~1.3 minutes). |
Since you have your own private compute node, you can use all its resources. It has 68 128 cores, so re-run the R2 alignment asking for 60 execution threads.
| Code Block |
|---|
bwa aln -t 60 sacCer3/sacCer3.fa fastq/Sample_Yeast_L005_R2.cat.fastq.gz > yeast_pe_R2.sai |
Exercise: How much of a speedup did you seen when aligning the R2 file with 20 60 threads?
| Expand |
|---|
|
The last few lines of bwa's execution output should look something like this: | Code Block |
|---|
| [bwa_aln] 17bp reads: max_diff = 2
[bwa_aln] 38bp reads: max_diff = 3
[bwa_aln] 64bp reads: max_diff = 4
[bwa_aln] 93bp reads: max_diff = 5
[bwa_aln] 124bp reads: max_diff = 6
[bwa_aln] 157bp reads: max_diff = 7
[bwa_aln] 190bp reads: max_diff = 8
[bwa_aln] 225bp reads: max_diff = 9
[bwa_aln_core] calculate SA coordinate... 266.70 sec
[bwa_aln_core] write to the disk... 0.04 sec
[bwa_aln_core] 262144 sequences have been processed.
[bwa_aln_core] calculate SA coordinate... 268.94 sec
[bwa_aln_core] write to the disk... 0.03 sec
[bwa_aln_core] 524288 sequences have been processed.
[bwa_aln_core] calculate SA coordinate... 72.26 sec
[bwa_aln_core] write to the disk... 0.01 sec
[bwa_aln_core] 592180 sequences have been processed.
[main] Version: 0.7.17-r1188
[main] CMD: /usr/local/bin/bwa aln -t 60 sacCer3/sacCer3.fa fastq/Sample_Yeast_L005_R2.cat.fastq.gz
[main] Real time: 195.872013 sec; CPU: 617142.095813 sec |
So the R2 alignment took only ~20 ~5 seconds (real time), or 615+ times as fast as with only one processing thread. Note, though, that the CPU time with 60 threads was greater (617 142.8 sec) than with only 1 thread (124 77.6 sec). That's because of the thread management overhead when using multiple threads. |
...
Here is the command line statement you need. Just execute it on the command line.
| Expand |
|---|
|
|
| title | Pairing of BWA R1 and R2 aligned reads |
|---|
bwa sampe sacCer3/sacCer3.fa yeast_R1.sai yeast_R2.sai \
fastq/Sample_Yeast_L005_R1.cat.fastq.gz \
fastq/Sample_Yeast_L005_R2.cat.fastq.gz > yeast_pairedend.sam |
You should now have a SAM file (yeast_pairedend.sam) that contains the alignments. It's just a text file, so take a look with head, more, less, tail, or whatever you feel like. Later you'll learn additional ways to analyze the data with samtools once you create a BAM file.
Exercise: What kind of information is in the first lines of the SAM file?
| Expand |
|---|
|
The SAM file has a number of header lines, which all start with an at sign ( @ ). The @SQ lines describe each contig (chromosome) and its length. There is also a @PG line that describes the way the bwa sampe was performed. |
Exercise: How many alignment records (not header records) are in the SAM file?
...
This looks for the pattern '^HWI' which is the start of every read name (which starts every alignment record).
Remember -c says just count the records, don't display them.
| Code Block |
|---|
|
grep -P -c '^HWI' yeast_pairedend.sam |
Or use the -v (invert) option to tell grep to print all lines that don't match a particular pattern; here, all header lines, which start with @.
| Code Block |
|---|
|
grep -P -v -c '^@' yeast_pairedend.sam |
# Copy the FASTA files for building references
mkdir -p $SCRATCH/core_ngs/references
cp $CORENGS/references/fasta/*.fa $SCRATCH/core_ngs/references/fasta/
# Copy a pre-built bwa index for sacCer3
mkdir -p $SCRATCH/core_ngs/references/bwa/sacCer3
cp $CORENGS/references/bwa/sacCer3/*.* $SCRATCH/core_ngs/references/bwa/sacCer3/
# Get the FASTQ to align
mkdir -p $SCRATCH/core_ngs/alignment/fastq
cp $CORENGS/alignment/*fastq.gz $SCRATCH/core_ngs/alignment/fastq/
# Stage the BWA .sai files
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
ln -sf ../fastq
ln -sf ../../references/bwa/sacCer3
cp $CORENGS/catchup/yeast_bwa/*.sai .
|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Pairing of BWA R1 and R2 aligned reads |
|---|
|
cd $SCRATCH/core_ngs/alignment/yeast_bwa
bwa sampe sacCer3/sacCer3.fa yeast_pe_R1.sai yeast_pe_R2.sai \
fastq/Sample_Yeast_L005_R1.cat.fastq.gz \
fastq/Sample_Yeast_L005_R2.cat.fastq.gz > yeast_pe.sam |
You should now have a SAM file (yeast_pe.sam) that contains the alignments.
Exercise: How many lines does the SAM file have? How does this compare to the number of input sequences (R1+R2)?
| Expand |
|---|
|
wc -l yeast_pe.sam reports 1,184,378 lines The alignment SAM file will contain records for both R1 and R2 reads, so we need to count sequences in both files. zcat ./fastq/Sample_Yeast_L005_R[12]*gz | wc -l | awk '{print $1/4}' reports 1,184,360 reads that were aligned So the SAM file has 18 more lines than the R1+R2 total. These are the header records that appear before any alignment records. |
It's just a text file, so take a look with head, more, less, tail, or whatever you feel like. Later you'll learn additional ways to analyze the data with samtools once you create a BAM file.
Exercise: What kind of information is in the first lines of the SAM file?
| Expand |
|---|
|
The SAM file has a number of header lines, which all start with an at sign ( @ ). The @SQ lines describe each contig (chromosome) and its length. There is also a @PG line that describes the way the bwa sampe was performed. |
Exercise: How many alignment records (not header records) are in the
| Expand |
|---|
|
| There are 1,184,360 alignment records. |
Exercise: How many sequences were in the R1 and R2 FASTQ files combined?
| Expand |
|---|
|
zcat fastq/Sample_Yeast_L005_R[12].cat.fastq.gz | wc -l | awk '{print $1/4}'
|
| Expand |
|---|
|
| There were a total of 1,184,360 original sequences (R1s + R2s) |
Exercises:
...
SAM file?
| Expand |
|---|
|
Both R1 and R2 reads must have separate alignment records, because there were 1,184,360 R1+R2 reads and the same number of alignment records. The SAM file must contain both mapped and un-mapped reads, because there were 1,184,360 R1+R2 reads and the same number of alignment records. Alignment records occur in the same read-name order as they did in the FASTQ, except that they come in pairs. The R1 read comes 1st, then the corresponding R2. This is called read name ordering. |
Using cut to isolate fields
Recall the format of a SAM alignment record:
 Image Removed
Image Removed
|
This looks for the pattern '^HWI' which is the start of every read name (which starts every alignment record).
Remember -c says just count the records, don't display them. | Code Block |
|---|
| grep -P -c '^HWI' yeast_pe.sam |
Or use the -v (invert) option to tell grep to print all lines that don't match a particular pattern; here, all header lines, which start with @. | Code Block |
|---|
| grep -P -v -c '^@' yeast_pe.sam |
|
| Expand |
|---|
|
| There are 1,184,360 alignment records. |
Exercises:
- Does the SAM file contain both mapped and un-mapped reads?
- What is the order of the alignment records in this SAM file?
| Expand |
|---|
|
The SAM file must contain both mapped and un-mapped reads, because there were 1,184,360 R1+R2 reads and the same number of alignment records. Alignment records occur in the same read-name order as they did in the FASTQ, except that they come in pairs. The R1 read comes 1st, then the corresponding R2. This is called read name ordering. |
Using cut to isolate fields
Recall the format of a SAM alignment record:
 Image Added
Image Added
Suppose you Suppose you wanted to look only at field 3 (contig name) values in the SAM file. You can do this with the handy cut command. Below is a simple example where you're asking cut to display the 3rd column value for the last 10 alignment records.
| Expand |
|---|
|
| Code Block |
|---|
| # Stage the aligned SAM file
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cp $CORENGS/catchup/yeast_bwa/yeast_pe.sam .
|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Cut syntax for a single field |
|---|
|
tail yeast_pairedendpe.sam | cut -f 3 |
By default cut assumes the field delimiter is Tab, which is the delimiter used in the majority of NGS file formats. You can specify a different delimiter with the -d option.
...
| Code Block |
|---|
| language | bash |
|---|
| title | Cut syntax for multiple fields |
|---|
|
tail -20 yeast_pairedendpe.sam | cut -f 2-6,9 |
You may have noticed that some alignment records contain contig names (e.g. chrV) in field 3 while others contain an asterisk ( * ). The * means the record didn't map. We're going to use this heuristic along with cut to see about how many records represent aligned sequences. (Note this is not the strictly correct method of finding unmapped reads because not all unmapped reads have an asterisk in field 3. Later you'll see how to properly distinguish between mapped and unmapped reads using samtools.)
...
| Code Block |
|---|
| language | bash |
|---|
| title | Grep pattern that doesn't match header |
|---|
|
# the ^@ pattern matches lines starting with @ (only header lines),
# and -v says output lines that don't match
grep -v -P '^@' yeast_pairedendpe.sam | head |
Ok, it looks like we're seeing only alignment records. Now let's pull out only field 3 using cut:
...
| Code Block |
|---|
| language | bash |
|---|
| title | Filter contig name of * (unaligned) |
|---|
|
grep -v -P '^@' yeast_pairedendpe.sam | cut -f 3 | grep -v '*' | head |
...
| Code Block |
|---|
| language | bash |
|---|
| title | Count aligned SAM records |
|---|
|
grep -v -P '^@' yeast_pairedendpe.sam | cut -f 3 | grep -v '*' | wc -l |
...
| Expand |
|---|
| title | Make sure you're in a idev session |
|---|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Start an idev session |
|---|
| idev -pm normal120 -mN 1201 -A UT-2015-05-18OTH21164 -r CoreNGS-Thu
# or
idev -m 90 -N 1 -n 68 A OTH21164 -p development |
|
| Code Block |
|---|
|
# If not already loaded
module load biocontainers # takes a while
module load samtools
samtools |
...
| Code Block |
|---|
| title | SAMtools suite usage |
|---|
|
Program: samtools (Tools for alignments in the SAM format)
Version: 1.109 (using htslib 1.109)
Usage: samtools <command> [options]
Commands:
-- Indexing
dict create a sequence dictionary file
faidx index/extract FASTA
fqidx index/extract FASTQ
index index alignment
-- Editing
calmd recalculate MD/NM tags and '=' bases
fixmate fix mate information
reheader replace BAM header
targetcut cut fosmid regions (for fosmid pool only)
addreplacerg adds or replaces RG tags
markdup mark duplicates
-- File operations
collate shuffle and group alignments by name
cat concatenate BAMs
merge merge sorted alignments
mpileup multi-way pileup
sort sort alignment file
split splits a file by read group
quickcheck quickly check if SAM/BAM/CRAM file appears intact
fastq converts a BAM to a FASTQ
fasta converts a BAM to a FASTA
-- Statistics
bedcov read depth per BED region
coverage alignment depth and percent coverage
depth compute the depth
flagstat simple stats
idxstats BAM index stats
phase phase heterozygotes
stats generate stats (former bamcheck)
-- Viewing
flags explain BAM flags
tview text alignment viewer
view SAM<->BAM<->CRAM conversion
depad convert padded BAM to unpadded BAM |
In this exercise, we will explore five utilities provided by samtools: view, sort, index, flagstat, and idxstats. Each of these is executed in one line for a given SAM/BAM file. In the SAMtools/BEDtools sections tomorrow we will explore samtools in capabilities more in depth.
| Warning |
|---|
| title | Know your samtools version! |
|---|
|
There are two main "eras" of SAMtools development: - "original" samtools
- v 0.1.19 is the last stable version
- "modern" samtools
- v 1.0, 1.1, 1.2 – avoid these (very buggy!)
- v 1.3+ – finally stable!
Unfortunately, some functions with the same name in both version eras have different options and arguments! So be sure you know which version you're using. (The samtools version is usually reported at the top of its usage listing). TACC BioContainers also offers the original samtools version: samtools/ctr-0.1.19--3. |
...
The samtools view utility provides a way of converting between SAM (text) and BAM (binary, compressed) format. It also provides many, many other functions which we will discuss lster. To get a preview, execute samtools view without any other arguments. You should see:
...
| Expand |
|---|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Get the alignment exercises files |
|---|
| mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cp $CORENGS/catchup/yeast_bwa/yeast_pairedendpe.sam .
|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Convert SAM to binary BAM |
|---|
|
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cat yeast_pairedend.sam | samtools view -b -o yeast_pe.sam > yeast_pairedendpe.bam |
- the -b option tells the tool to output BAM format
- the -o option specifies the name of the output BAM file that will be created
- we pipe the entire SAM file to samtools view so that the header records are included (required for SAM → BAM conversion)
- samtools view reads its input from standard input by default
How do you look at the BAM file contents now? That's simple. The BAM file is a binary file, not a text file, so how do you look at its contents now? Just use samtools view without the -b option. Remember to pipe output to a pager!
| Code Block |
|---|
| language | bash |
|---|
| title | View BAM records |
|---|
|
samtools view yeast_pairedendpe.bam | more
|
Notice that this does not show us the header record we saw at the start of the SAM file.
Exercise: What samtools view option will include the header records in its output? Which option would show only the header records?
| Expand |
|---|
|
samtools view | less
then search for "header" ( /header ) |
| Expand |
|---|
|
samtools view -h shows header records along with alignment records. samtools view -H shows header records only. |
...
Looking at some of the alignment record information (e.g. samtools view yeast_pairedendpe.bam | cut -f 1-4 | more), you will notice that read names appear in adjacent pairs (for the R1 and R2), in the same order they appeared in the original FASTQ file. Since that means the corresponding mappings are in no particular order, searching through the file very inefficient. samtools sort re-orders entries in the SAM file either by locus (contig name + coordinate position) or by read name.
...
| Code Block |
|---|
|
Usage: samtools sort [options...] [in.bam]
Options:
-l INT Set compression level, from 0 (uncompressed) to 9 (best)
-m INT Set maximum memory per thread; suffix K/M/G recognized [768M]
-n Sort by read name
-t TAG Sort by value of TAG. Uses position as secondary index (or read name if -n is set)
-o FILE Write final output to FILE rather than standard output
-T PREFIX Write temporary files to PREFIX.nnnn.bam
--no-PG do not add a PG line
-- --input-fmt-option OPT[=VAL]
Specify a single input file format option in the form
of OPTION or OPTION=VALUE
-O, --output-fmt FORMAT[,OPT[=VAL]]...
Specify output format (SAM, BAM, CRAM)
--output-fmt-option OPT[=VAL]
Specify a single output file format option in the form
of OPTION or OPTION=VALUE
--reference FILE
Reference sequence FASTA FILE [null]
-@, --threads INT
Number of additional threads to use [0]
--verbosity INT
Set level of verbosity |
In most cases you will be sorting a BAM file from name order to locus order. You can use either -o or redirection with > to control the output.
| Expand |
|---|
|
Copy aligned yeast BAM file | Code Block |
|---|
| # Stage the aligned yeast SAM and BAM files
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cp $CORENGS/catchup/yeast_bwa/yeast_pairedend.bampe.[bs]am . |
|
To sort the paired-end yeast BAM file by position, and get a BAM file named yeast_pairedendpe.sort.bam as output, execute the following command:
| Code Block |
|---|
| language | bash |
|---|
| title | Sort a BAM file |
|---|
|
cd $SCRATCH/core_ngs/alignment/yeast_bwa
samtools sort -O bam -T yeast_pairedendpe.tmp yeast_pairedendpe.bam > yeast_pairedendpe.sort.bam |
- The -O options says the Output format should be BAM
- The -T options gives a prefix for Temporary files produced during sorting
- sorting large BAMs will produce many temporary files during processing
- make sure the temporary file prefix is different from the input BAM file prefix!
- By default sort writes its output to standard output, so we use > to redirect to a file named yeast_pairedend.sort.bam
Exercise: Compare the file sizes of the yeast_pariedend pe .sam, .bam, and .sort.bam files and explain why they are different.
| Expand |
|---|
|
| Code Block |
|---|
| ls -lh yeast_pairedendpe.* |
|
| Expand |
|---|
|
The yeast_pairedendpe.sam text file is the largest at ~348 MB .because it is an uncompressed text file. The name-ordered binary yeast_pairedendpe.bam text file only about 1/3 that size, ~110 ~111 MB. They contain exactly the same records, in the same order, but conversion from text to binary results in a much smaller file. The coordinate-ordered binary yeast_pairedendpe.sort.bam file is even slightly smaller, ~91 ~92 MB. This is because BAM files are actually customized gzip-format files. The customization allows blocks of data (e.g. all alignment records for a contig) to be represented in an even more compact form. You can read more about this in section 4 of the SAM format specification. |
...
| Code Block |
|---|
| language | bash |
|---|
| title | Index a sorted bam |
|---|
|
samtools index yeast_pairedendpe.sort.bam |
This will produce a file named yeast_pairedendpe.bam.bai.
Most of the time when an index is required, it will be automatically located as long as it is in the same directory as its BAM file and shares the same name up until the .bai extension.
...
| Expand |
|---|
|
| Code Block |
|---|
| ls -lh yeast_pairedendpe.sort.bam* |
|
| Expand |
|---|
|
While the yeast_pairedendpe.sort.bam text file is ~91 ~92 MB, its index (yeast_pairedendpe.sort.bai) is only 20 KB. |
samtools flagstat
...
Here's how to run samtools flagstat and both see the output in the terminal and save it in a file – the samtools flagstat standard output is piped to tee, which both writes it to the specified file and sends it to its standard output:
| Code Block |
|---|
| language | bash |
|---|
| title | Run samtools flagstat using tee |
|---|
|
samtools flagstat yeast_pairedend.sort.bam | tee yeast_pariedend.flagstat.txt |
You should see something like this:
| Code Block |
|---|
| title | samtools flagstat output |
|---|
|
1184360 + 0 in total (QC-passed reads + QC-failed reads)
0 + 0 secondary
0 + 0 supplementary
0 + 0 duplicates
547664 + 0 mapped (46.24% : N/A)
1184360 + 0 paired in sequencing
592180 + 0 read1
592180 + 0 read2
473114 + 0 properly paired (39.95% : N/A)
482360 + 0 with itself and mate mapped
65304 + 0 singletons (5.51% : N/A)
534 + 0 with mate mapped to a different chr
227 + 0 with mate mapped to a different chr (mapQ>=5) |
Ignore the "+ 0" addition to each line - that is a carry-over convention for counting QA-failed reads that is no longer relevant.
The most important statistic is the mapping rate (here 46%) but this readout also allows you to verify that some common expectations (e.g. that about the same number of R1 and R2 reads aligned, and that most mapped reads are proper pairs) are met.
Exercise: What proportion of mapped reads were properly paired?
| Expand |
|---|
|
Divide the number of properly paired reads by the number of mapped reads: | Code Block |
|---|
| awk 'BEGIN{ print 473114 / 547664 }'
# or
echo $(( 473114 * 100 / 547664 ))
|
|
| Expand |
|---|
|
About 86% of mapped read were properly paired. This is actually a bit on the low side for ChIP-seq alignments which typically over 90%. |
samtools idxstats
More information about the alignment is provided by the samtools idxstats report, which shows how many reads aligned to each contig in your reference. Note that samtools idxstats must be run on a sorted, indexed BAM file.
| Code Block |
|---|
| language | bash |
|---|
| title | Use samtools idxstats to summarize mapped reads by contig |
|---|
|
samtools idxstats yeast_pairedend.sort.bam | tee yeast_pairedend.idxstats.txt |
| Code Block |
|---|
| language | bash |
|---|
| title | samtools idxstats output |
|---|
|
chrI 230218 8820 1640
chrII 813184 36616 4026
chrIII 316620 13973 1530
chrIV 1531933 72675 8039
chrV 576874 27466 2806
chrVI 270161 10866 1222
chrVII 1090940 50893 5786
chrVIII 562643 24672 3273
chrIX 439888 16246 1739
chrX 745751 31748 3611
chrXI 666816 28017 2776
chrXII 1078177 54783 10124
chrXIII 924431 40921 4556
chrXIV 784333 33070 3703
chrXV 1091291 48714 5150
chrXVI 948066 44916 5032
chrM 85779 3268 291
* 0 0 571392 |
The output has four tab-delimited columns:
- contig name
- contig length
- number of mapped reads
- number of unmapped reads
The reason that the "unmapped reads" field for named chromosomes is not zero is that the aligner may initially assign a potential mapping (contig name and start coordinate) to a read, but then mark it later as unampped if it does meet various quality thresholds.
| Tip |
|---|
If you're mapping to a non-genomic reference such as miRBase miRNAs or another set of genes (a transcriptome), samtools idxstats gives you a quick look at quantitative alignment results. |
Exercise #3: PE alignment with BioITeam scripts
Now that you've done everything the hard way, let's see how to do run an alignment pipeline using a BWA alignment script maintained by the BioITeam, /work2/projects/BioITeam/common/script/align_bwa_illumina.sh. Type in the script name to see its usage.
| Code Block |
|---|
|
align_bwa_illumina.sh 2021_06_05
Align Illumina SE or PE data with bwa. Produces a sorted, indexed,
duplicate-marked BAM file and various statistics files. Usage:
align_bwa_illumina.sh <aln_mode> <in_file> <out_pfx> <assembly> [ paired trim_sz trim_sz2 seq_fmt qual_fmt ]
Required arguments:
aln_mode Alignment mode, either global (bwa aln) or local (bwa mem).
in_file For single-end alignments, path to input sequence file.
For paired-end alignments using fastq, path to the the R1
fastq file which must contain the string 'R1' in its name.
The corresponding 'R2' must have the same path except for 'R1'.
out_pfx Desired prefix of output files in the current directory.
assembly One of hg38, hg19, hg38, mm10, mm9, sacCer3, sacCer1, ce11, ce10,
danRer7, hs_mirbase, mm_mirbase, or reference index prefix.
Optional arguments:
paired 0 = single end alignment (default); 1 = paired end.
trim_sz Size to trim reads to. Default 0 (no trimming)
trim_sz2 Size to trim R2 reads to for paired end alignments.
Defaults to trim_sz
seq_fmt Format of sequence file (fastq, bam or scarf). Default is
fastq if the input file has a '.fastq' extension; scarf
if it has a '.sequence.txt' extension.
qual_type Type of read quality scores (sanger, illumina or solexa).
Default is sanger for fastq, illumina for scarf.
Environment variables:
show_only 1 = only show what would be done (default not set)
aln_args other bowtie2 options (e.g. '-T 20' for mem, '-l 20' for aln)
no_markdup 1 = don't mark duplicates (default 0, mark duplicates)
run_fastqc 1 = run fastqc (default 0, don't run). Note that output
will be in the directory containing the fastq files.
keep 1 = keep unsorted BAM (default 0, don't keep)
bwa_bin BWA binary to use. Default bwa 0.7.x. Note that bwa 0.6.2
or earlier should be used for scarf and other short reads.
also: NUM_THREADS, BAM_SORT_MEM, SORT_THREADS, JAVA_MEM_ARG
Examples:
align_bwa_illumina.sh local ABC_L001_R1.fastq.gz my_abc hg38 1
align_bwa_illumina.sh global ABC_L001_R1.fastq.gz my_abc hg38 1 50
align_bwa_illumina.sh global sequence.txt old sacCer3 0 '' '' scarf solexa |
There are lots of bells and whistles in the arguments, but the most important are the first few:
- aln_mode – whether to perform a global or local alignment (the 1st argument must be one of those words)
- global mode uses the bwa aln workflow as we did above
- local mode uses the bwa mem command
- in_file – full or relative path to the FASTQ file (just the R1 fastq if paired end). Can be compressed (.gz)
- out_pfx – prefix for all the output files produced by the script. Should relate back to what the data is.
- assembly – genome assembly to use.
- there are pre-built indexes for some common eukaryotes (hg38, hg19, mm10, mm9, danRer7, sacCer3) that you can use
- or provide a full path for a bwa reference index you have built somewhere
- paired flag – 0 means single end (the default); 1 means paired end
- trim_sz – if you want the FASTQ hard trimmed down to a specific length before alignment, supply that number here
We're going to run this script and a similar Bowtie2 alignment script, on the yeast data using the TACC batch system. In a new directory, copy over the commands and submit the batch job. We ask for 2 hours (-t 02:00:00) with 4 tasks/node (-w 4); since we have 4 commands, this will run on 1 compute node.
| Expand |
|---|
| title | Catch up (if needed) |
|---|
|
| Code Block |
|---|
| # Copy over the Yeast data if needed
mkdir -p $SCRATCH/core_ngs/alignment/fastq
cp $CORENGS/alignment/Sample_Yeast*.gz $SCRATCH/core_ngs/alignment/fastq/
|
|
| Code Block |
|---|
| language | bash |
|---|
| title | Run multiple alignments using the TACC batch system |
|---|
|
# Make sure you're not in an idev session by looking at the hostname
hostname
# If the hostname looks like "c455-004.stampede2.tacc.utexas.edu", exit the idev session
# Make a new alignment directory for running these scripts
mkdir -p $SCRATCH/core_ngs/alignment/bwa_script
cd $SCRATCH/core_ngs/alignment/bwa_script
ln -s -f ../fastq
# Copy the alignment commands file and submit the batch job
cp $CORENGS/tacc/aln_script.cmds .
launcher_creator.py -j aln_script.cmds -n aln_script -t 02:00:00 -w 4 -a UT-2015-05-18 -q normal
sbatch --reservation=BIO_DATA_week_1 aln_script.slurm
showq -u |
While we're waiting for the job to complete, lets look at the aln_script.cmds file.
| Code Block |
|---|
| language | bash |
|---|
| title | Commands to run multiple alignment scripts |
|---|
|
/work2/projects/BioITeam/common/script/align_bwa_illumina.sh global ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz bwa_global sacCer3 1 50
/work2/projects/BioITeam/common/script/align_bwa_illumina.sh local ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz bwa_local sacCer3 1
/work2/projects/BioITeam/common/script/align_bowtie2_illumina.sh global ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz bt2_global sacCer3 1 50
/work2/projects/BioITeam/common/script/align_bowtie2_illumina.sh local ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz bt2_local sacCer3 1 |
Notes:
- The 1st command performs a paired-end BWA global alignment (similar to above), but asks that the 100 bp reads be trimmed to 50 first.
- we refer to the pre-built index for yeast by name: sacCer3
- this index is located in the /work/projects/BioITeam/ref_genome/bwa/bwtsw/sacCer3/ directory
- we provide the name of the R1 FASTQ file
- because we request a PE alignment (the 1 argument) the script will look for a similarly-named R2 file.
- all output files associated with this command will be named with the prefix bwa_global.
- The 2nd command performs a paired-end BWA local alignment.
- all output files associated with this command will be named with the prefix bwa_local.
- no trimming is requested because the local alignment should ignore 5' and 3' bases that don't match the reference genome
- The 3rd command performs a paired-end Bowtie2 global alignment.
- the Bowtie2 alignment script has the same first arguments as the BWA alignment script.
- all output files associated with this command will be named with the prefix bt2_global.
- again, we specify that reads should first be trimmed to 50 bp.
- The 4th command performs a paired-end Bowtie2 local alignment.
- all output files associated with this command will be named with the prefix bt2_local.
- again, no trimming is requested for the local alignment.
Output files
This alignment pipeline script performs the following steps:
- Hard trims FASTQ, if optionally specified (fastx_trimmer)
- Performs the global or local alignment (here, a PE alignment)
- BWA global: bwa aln the R1 and R2 separately, then bwa sampe to produce a SAM file
- BWA local: call bwa mem with both R1 and R2 to produce a SAM file
- Bowtie2 global: call bowtie2 --global with both R1 and R2 to produce a SAM file
- Bowtie2 local: call bowtie2 --local with both R1 and R2 to produce a SAM file
- Converts SAM to BAM (samtools view)
- Sorts the BAM (samtools sort)
- Marks duplicates (Picard MarkDuplicates)
- Indexes the sorted, duplicate-marked BAM (samtools index)
- Gathers statistics (samtools idxstats, samtools flagstat, plus a custom statistics script of Anna's)
- Removes intermediate files
There are a number of output files, with the most important being those desribed below.
- <prefix>.align.log – Log file of the entire alignment process.
- check the tail of this file to make sure the alignment was successful
- <prefix>.sort.dup.bam – Sorted, duplicate-marked alignment file.
- <prefix>.sort.dup.bam.bai – Index for the sorted, duplicate-marked alignment file
- <prefix>.flagstat.txt – samtools flagstat output
- <prefix>.idxstats.txt – samtools idxstats output
- <prefix>.samstats.txt – Summary alignment statistics from Anna's stats script
- <prefix>.iszinfo.txt – Insert size statistics (for paired-end alignments) from Anna's stats script
Verifying alignment success
The alignment log will have a "I ran successfully" message at the end if all went well, and if there was an error, the important information should also be at the end of the log file. So you can use tail to check the status of an alignment. For example:
| Code Block |
|---|
| language | bash |
|---|
| title | Checking the alignment log file |
|---|
|
tail bwa_global.align.log |
This will show something like:
| Code Block |
|---|
..Done alignmentUtils.pl bamstats - 2020-06-14 23:19:38
.. samstats file 'bwa_global.samstats.txt' exists and is not empty - 2020-06-14 23:19:38
===============================================================================
## Cleaning up files (keep 0) - 2020-06-14 23:19:38
===============================================================================
ckRes 0 cleanup
===============================================================================
## All bwa alignment tasks completed successfully! - 2020-06-14 23:19:38
=============================================================================== |
Notice that success message: "All bwa alignment tasks completed successfully!". It should only appear once in any successful alignment log.
When multiple alignment commands are run in parallel it is important to check them all, and you can use grep looking for part of the unique success message to do this. For example:
| Code Block |
|---|
| language | bash |
|---|
| title | Count the number of successful alignments |
|---|
|
grep 'completed successfully!' *align.log | wc -l |
If this command returns 4 (the number of alignment tasks we performed), all went well, and we're done.
But what if something went wrong? How can we tell which alignment task was not successful? You could tail the log files one by one to see which one(s) don't have the message, but you can also use a special grep option to do this work.
| Code Block |
|---|
| language | bash |
|---|
| title | Check for failed alignment tasks |
|---|
|
grep -L 'completed successfully' *.align.log |
The -L option tells grep to only print the filenames that don't contain the pattern. Perfect! To see happens in the case of failure, try it on a file that doesn't contain that message:
| Code Block |
|---|
|
grep -L 'completed successfully' aln_script.cmds |
Checking alignment statistics
The <prefix>.samstats.txt statistics files produced by the alignment pipeline has a lot of good information in one place. If you look at bwa_global.samstats.txt you'll see something like this:
| Code Block |
|---|
| title | <prefix>.samstats.txt output |
|---|
|
-----------------------------------------------
Aligner: bwa
Total sequences: 1184360
Total mapped: 539079 (45.5 %)
Total unmapped: 645281 (54.5 %)
Primary: 539079 (100.0 %)
Secondary:
Duplicates: 249655 (46.3 %)
Fwd strand: 267978 (49.7 %)
Rev strand: 271101 (50.3 %)
Unique hit: 503629 (93.4 %)
Multi hit: 35450 (6.6 %)
Soft clip:
All match: 531746 (98.6 %)
Indels: 7333 (1.4 %)
Spliced:
-----------------------------------------------
Total PE seqs: 1184360
PE seqs mapped: 539079 (45.5 %)
Num PE pairs: 592180
F5 1st end mapped: 372121 (62.8 %)
F3 2nd end mapped: 166958 (28.2 %)
PE pairs mapped: 80975 (13.7 %)
PE proper pairs: 16817 (2.8 %)
----------------------------------------------- |
Since this was a paired end alignment there is paired-end specific information reported.
You can also view statistics on insert sizes for properly paired reads in the bwa_global.iszinfo.txt file. This tells you the average (mean) insert size, standard deviation, mode (most common value), and fivenum values (minimum, 1st quartile, median, 3rd quartile, maximum).
| Code Block |
|---|
| title | <prefix>.iszinfo.txt output |
|---|
|
Insert size stats for: bwa_global
Number of pairs: 16807 (proper)
Number of insert sizes: 406
Mean [-/+ 1 SD]: 296 [176 416] (sd 120)
Mode [Fivenum]: 228 [51 224 232 241 500] |
A quick way to check alignment stats if you have run multiple alignments is again to use grep. For example:
| Code Block |
|---|
| language | bash |
|---|
| title | Review multiple alignment rates |
|---|
|
grep 'Total mapped' *samstats.txt |
will produce output like this:
| Code Block |
|---|
bt2_global.samstats.txt: Total mapped: 602893 (50.9 %)
bt2_local.samstats.txt: Total mapped: 788069 (66.5 %)
bwa_global.samstats.txt: Total mapped: 539079 (45.5 %)
bwa_local.samstats.txt: Total mapped: 1008000 (76.5 % |
Exercise: How would you list the median insert size for all the alignments?
| Expand |
|---|
|
That information is in the *.iszinfo.txt files, on the line labeled Mode. The median value is th 3rd value in the 5 fivnum values; it is the 7th whitespace-separated field on the Mode line.
|
| Expand |
|---|
|
Use grep to isolate the Mode line, and awk to isolate the median value field: | Code Block |
|---|
| grep 'Mode' *.iszinfo.txt | awk '{print $1,"Median insert size:",$7}' |
|
TACC batch system considerations
The great thing about pipeline scripts like this is that you can perform alignments on many datasets in parallel at TACC, and they are written to take advantage of having multiple cores on TACC nodes where possible.
On the stampede2, with its 68 physical cores per node, they are designed to run best with no more than 4 tasks per node.
sends it to its standard output:
| Expand |
|---|
|
| Code Block |
|---|
| # Stage the aligned yeast SAM and BAM files
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cp $CORENGS/catchup/yeast_bwa/yeast_pe.sort.bam* . |
|
| Code Block |
|---|
| language | bash |
|---|
| title | Run samtools flagstat using tee |
|---|
|
samtools flagstat yeast_pe.sort.bam | tee yeast_pe.flagstat.txt |
You should see something like this:
| Code Block |
|---|
| title | samtools flagstat output |
|---|
|
1184360 + 0 in total (QC-passed reads + QC-failed reads)
0 + 0 secondary
0 + 0 supplementary
0 + 0 duplicates
547664 + 0 mapped (46.24% : N/A)
1184360 + 0 paired in sequencing
592180 + 0 read1
592180 + 0 read2
473114 + 0 properly paired (39.95% : N/A)
482360 + 0 with itself and mate mapped
65304 + 0 singletons (5.51% : N/A)
534 + 0 with mate mapped to a different chr
227 + 0 with mate mapped to a different chr (mapQ>=5) |
Ignore the "+ 0" addition to each line - that is a carry-over convention for counting "QA-failed reads" that is no longer relevant.
The most important statistic is the mapping rate (here 46%) but this readout also allows you to verify that some common expectations (e.g. that about the same number of R1 and R2 reads aligned, and that most mapped reads are proper pairs) are met.
Exercise: What proportion of mapped reads were properly paired?
| Expand |
|---|
|
Divide the number of properly paired reads by the number of mapped reads: | Code Block |
|---|
| awk 'BEGIN{ print 473114 / 547664 }'
# or
echo $(( 473114 * 100 / 547664 ))
# or
echo "473114 547664" | awk '{printf("%0.1f%%\n", 100*$1/$2)}'
|
|
| Expand |
|---|
|
About 86% of mapped read were properly paired. This is actually a bit on the low side for ChIP-seq alignments which typically over 90%. |
samtools idxstats
More information about the alignment is provided by the samtools idxstats report, which shows how many reads aligned to each contig in your reference. Note that samtools idxstats must be run on a sorted, indexed BAM file.
| Expand |
|---|
|
| Code Block |
|---|
| # Stage the aligned yeast SAM and BAM files
mkdir -p $SCRATCH/core_ngs/alignment/yeast_bwa
cd $SCRATCH/core_ngs/alignment/yeast_bwa
cp $CORENGS/catchup/yeast_bwa/yeast_pe.sort.bam* . |
|
| Code Block |
|---|
| language | bash |
|---|
| title | Use samtools idxstats to summarize mapped reads by contig |
|---|
|
samtools idxstats yeast_pe.sort.bam | tee yeast_pe.idxstats.txt |
Here we use the tee command which reports its standard input to standard output before also writing it to the specified file.
| Code Block |
|---|
| language | bash |
|---|
| title | samtools idxstats output |
|---|
|
chrI 230218 8820 1640
chrII 813184 36616 4026
chrIII 316620 13973 1530
chrIV 1531933 72675 8039
chrV 576874 27466 2806
chrVI 270161 10866 1222
chrVII 1090940 50893 5786
chrVIII 562643 24672 3273
chrIX 439888 16246 1739
chrX 745751 31748 3611
chrXI 666816 28017 2776
chrXII 1078177 54783 10124
chrXIII 924431 40921 4556
chrXIV 784333 33070 3703
chrXV 1091291 48714 5150
chrXVI 948066 44916 5032
chrM 85779 3268 291
* 0 0 571392 |
The output has four tab-delimited columns:
- contig name
- contig length
- number of mapped reads
- number of unmapped reads
The reason that the "unmapped reads" field for named chromosomes is not zero is that the aligner may initially assign a potential mapping (contig name and start coordinate) to a read, but then mark it later as unampped if it does meet various quality thresholds.
| Tip |
|---|
If you're mapping to a non-genomic reference such as miRBase miRNAs or another set of genes (a transcriptome), samtools idxstats gives you a quick look at quantitative alignment results |
| Tip |
|---|
| title | Always specify wayness 4 for alignment pipeline scripts |
|---|
|
These alignment scripts should always be run with a wayness of 4 (-w 4) in the stampede2 batch system, meaning at most 4 commands per node. |


