...
All GO terms have an ID that looks like GO:0006260 and a name like DNA replication.
All GO terms have alist of genes that belong to that particular term.
GO terms are hierarchical consisting of broader parent GO terms and narrower child GO terms. For example, DNA replication is a child of GO term:cellular metabolic process. DNA replication has child GO terms like regulation of DNA replication, strand elongation.
WHAT IS GO ENRICHMENT?
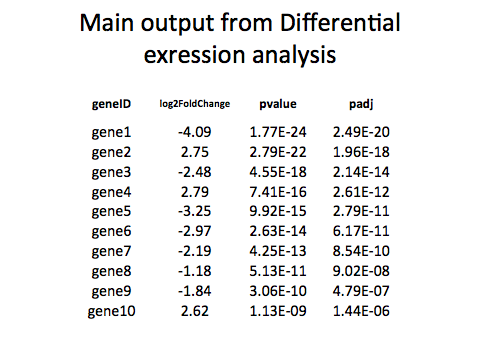
GO enrichment is a way of summarizing the FUNCTIONS AND TYPES of genes that are differentially expressed.
...
INPUT FILE 2: ALL (contains allall 14869 genes)
FBgn0000370
FBgn0025682
...
SCP THE DATA OVER TO YOUR COMPUTER:
| Code Block | ||
|---|---|---|
| ||
#ON STAMPEDE2LS5: copy the path for the ALL and DEG files pwd #ON LOCAL COMPUTER: from a terminal tab scp <username>@stampede2<username>@ls5.tacc.utexas.edu:<pathtofilesonstampede<pathtofileson/DEG> . scp <username>@stampede2<username>@ls5.tacc.utexas.edu:<pathtofilesonstampede<pathtofileson/ALL> . |
RUN GORILLA USING THE UNRANKED METHOD: http://cbl-gorilla.cs.technion.ac.il/
...
INPUT FILE: ALLRANKED (all genes, ranked by adjusted pvalue)
FBgn0000370
FBgn0025682
...
| Code Block | ||
|---|---|---|
| ||
##Command to pull out ALL gene ids, sorted by pvalueadjpvalue store it in a file called ALLRANKED #Remember we already sorted our results by adjusted pvalue in the deseq2 script before writing it out to a file. So you just #need to pull out the gene ids in the order it already is in. |
SCP THE DATA OVER TO YOUR COMPUTER:
| Code Block | ||
|---|---|---|
| ||
#ON STAMPEDE2LS5: copy the path for the ALL and DEG files pwd #ON LOCAL COMPUTER: from a terminal tab scp <username>@stampede2<username>@ls5.tacc.utexas.edu:<pathtofilesonstampede/DEG> . scp <username>@stampede2.tacc.utexas.edu:<pathtofilesonstampede/ALL><pathtofileson/ALLRANKED> . |
RUN GORILLA USING THE RANKED METHOD: http://cbl-gorilla.cs.technion.ac.il/