...
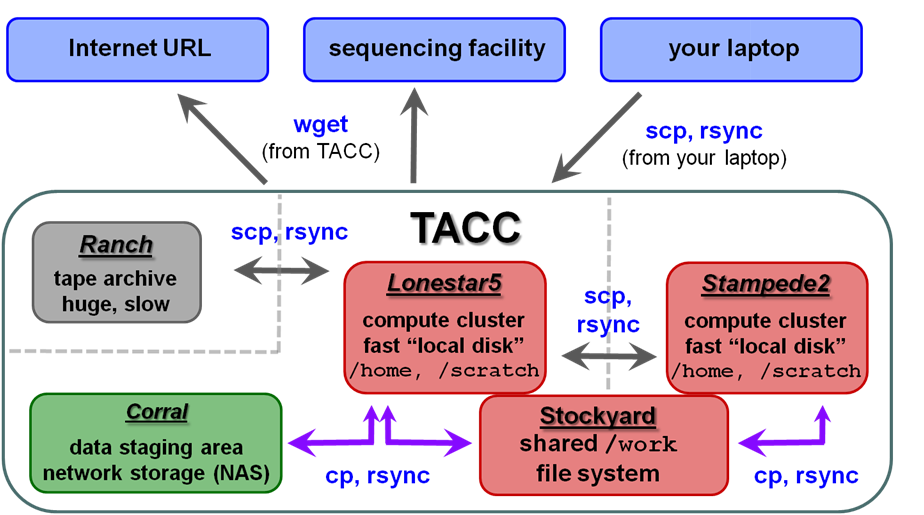
| TACC storage areas and Linux commands to access data (all commands to be executed at TACC except laptop-to-TACC copies, which must be executed on your laptop) |
Local file systems
There are 3 local file systems available on any TACC compute cluster (Lonestar5stampede2, stampede2 lonestar5, etc.), each with different characteristics. All these local file systems are very fast and set up for parallel I/O (Lustre file system).
On ls5 stampede2 these local file systems have the following characteristics:
| Home | WorkWork2 | Scratch | |
|---|---|---|---|
| quota | 10 GB | 1024 GB = 1 TB | 122+ PB (basically infinite) |
| policy | backed up | not backed up, not purged | not backed up, purged if not accessed recently (~10 days) |
| access command | cd | cdwcdw2 | cds |
| environment variable | $HOME | $STOCKYARD (root of the shared Work file system) $WORK $WORK2 (different sub-directory for each cluster) | $SCRATCH |
| root file system | /home | /workwork2 | /scratch |
| use for | Small files such as scripts that you don't want to lose. | Medium-sized artifacts you don't want to copy over all the time. For example, custom programs you install (these can get large), or annotation file used for analysis. | Large files accessed from batch jobs. Your starting files will be copied here from somewhere else, and your final results files will be copied elsewhere (e.g. stockyard, corral, or your BRCF POD). |
...
| Code Block |
|---|
--------------------- Project balances for user abattenh ---------------------- | Name Avail SUs Expires | Name Avail SUs Expires | | CancerGeneticsgenomeAnalysis 4856673 20182021-0903-3031 | A-cm10BioinformaticsResour 100 1096 2018-12-312020-06-30 | | UT-2015-05-18 21001000 20192021-03-31 | genomeAnalysisDNAdenovo 2500 2019-03-31 4969 2021-03-31 | | CancerGenetics 4856 2020-09-30 | A-cm10 8867 2020-12-31 | ------------------------ Disk quotas for user abattenh ------------------------- | Disk Usage (GB) Limit %Used File Usage Limit %Used | | /home1 0.0 10.0 0.1210 91153 1000000 0.0102 | | /workwork2 538614.5 1024.0 5260.5901 6105361094 3000000 2.04 | | /scratch 37252676.96 0.0 0.00 413732442 0 0.00 | ------------------------------------------------------------------------------- |
...
When you first login, you start in your home directory. Use these the cd, cdw2 and cds commands to change to your other file systems. Notice how your command prompt helpfully changes to show your location.
| Code Block | ||||
|---|---|---|---|---|
| ||||
cdwcdw2 cds cd |
| Tip |
|---|
The cd (change directory) command with no arguments takes you to your home directory on any Linux/Unix system. The cdw cdw2 and cds commands are specific to the TACC environment. |
...
TACC compute clusters now share a common Work file system called stockyard. So files in your Work area do not have to be copied, for example from ls5 to stampede2 to ls5 – they can be accessed directly from either cluster.
...
- $STOCKYARD - This refers to the root of your shared Work area
- e.g. /work/01063/abattenh (should be changed to /work2/01063/abattenh soon)
- e.g. /work/01063/abattenh (should be changed to /work2/01063/abattenh soon)
- $WORK or $WORK2 - Refers to a sub-directory of the shared Work area that is different for different clusters, e.g.:
- /work/01063/abattenh/lonestar on ls5
- /workwork2/01063/abattenh/stampede2 on stampede2
A mechanism for purchasing larger stockyard allocations (above the 1 TB basic quota) from TACC are in development.
The UT Austin BioInformatics Team, a loose group of bioinformatics researchers, maintains a common directory area on stockyard.
| Code Block | ||||
|---|---|---|---|---|
| ||||
ls /workwork2/projects/BioITeam |
Files we will use in this course are in a sub-directory there. The $CORENGS environment variable set in your login profile refers to this path.
| Code Block | ||||
|---|---|---|---|---|
| ||||
echo $CORENGS ls /workwork2/projects/BioITeam/projects/courses/Core_NGS_Tools |
...
corral is a gigantic (multiple PB) storage system (spinning disk) where researchers can store data. UT researchers may request up to 5 TB of corral storage through the normal TACC allocation request process. Additional space on corral can be rented for ~$85~$80/TB/year.
The UT/Austin BioInformatics Team also has an older, common directory area on corral.
| Code Block | ||||
|---|---|---|---|---|
| ||||
ls /corral-repl/utexas/BioITeam |
A couple of things A couple of things to keep in mind regarding corral:
...
ranch is a gigantic (multiple PB) tape archive system where researchers can archive data. All TACC users have an automatice 2 TB ranch allocation. UT researchers may request large larger (multi-TB) ranch storage allocations through the normal TACC allocation request process.
There is currently no charge for ranch storage. However, since the data is stored on tape it is not immediately available – robots find and mount appropriate tapes when the data is requested, and it can take minutes to hours for the data to appear on disk. ( The metadata about your data – the directory structures and file names – is always accessible, but the actual data in the files is not on disk until "staged". See the ranch user guide for more information: https://www.tacc.utexas.edu/user-services/user-guides/ranch-user-guide.
Once that data is staged to the ranch disk it can be copied to other places. However, the ranch file system is not mounted as a local file system from the stampede2 or ls5 clusters. So remote copy commands are always needed to copy data to and from ranch (e.g. scp, rsync).
...
The first task is to get this sequencing data to a permanent storage area. This should not be your laptop or one of the TACC local file systems! corral (or stockyard) is a great place for it, or a server maintained by your lab or company.We're going
to pretend – just for the sake of this class – that Here's an example of a "best practice". Wherever your permanent storage area is in your TACC work area. Execute these commands to make your "archive" directory and some sub-directories.
| Code Block | ||||
|---|---|---|---|---|
| ||||
mkdir -p $WORK/archive/original/2018_05.core_ngs |
Here's an example of a "best practice". Wherever your permanent storage area is, it should have a rational sub-directory structure that reflects its contents. It's easy to process a few NGS datasets, but when they start multiplying like tribbles, good organization and naming conventions will be the only thing standing between you and utter chaos!
For example:
, it should have a rational sub-directory structure that reflects its contents. It's easy to process a few NGS datasets, but when they start multiplying like tribbles, good organization and naming conventions will be the only thing standing between you and utter chaos!
For example:
- original – for original sequencing data (compressed FASTQ files)
- sub-directories named, for example, by year_month.<project_name>
- aligned – for alignment artifacts (BAM files, etc)
- sub-directories named, e.g., by
- sub-directories named, for example, by year_month.<project_name>
- aligned – for alignment artifacts (BAM files, etc)
- sub-directories named, e.g., by year_month.<project_name>
- analysis – further downstream analysis
- reasonably
- reasonably named sub-directories, often by project
- genome refs – reference genomes and other annotation files used in alignment and analysis
- sub-directories for different reference genomes
- e.g. ucsc/hg19, ucsc/sacCer3, mirbase/v20
- code – for scripts and programs you and others in your organization write
- ideally maintained in a version control system such as git, subversion or cvs.
- easiest to name sub-directories for people.
...
Get ready to run wget from the directory where you want to put the data.
Don't press Enter after the wget command – just put a space after it.
| Code Block | ||||
|---|---|---|---|---|
| ||||
mkdir -p $SCRATCH/archive/original/2021.core_ngs cd $WORK$SCRATCH/archive/original/2018_052021.core_ngs wget |
Here are two web links:
- httphttps://web.corral.tacc.utexas.edu/BioITeamBioinformaticsResource/CoreNGS/yeast_stuff/Sample_Yeast_L005_R1.cat.fastq.gzhttp
- https://web.corral.tacc.utexas.edu/BioITeamBioinformaticsResource/CoreNGS/yeast_stuff/Sample_Yeast_L005_R2.cat.fastq.gz
Right-click (Windows) or Control+click (Mac) on the 1st link in your browser, then select "Copy link location" from the menu. Now go back to your Terminal. Put your cursor after the space following the wget command then either right-click (Windows), or Paste (Command-V on Mac, Control-V on Windows). The command line to be executed should now look like this:
| Code Block | ||||
|---|---|---|---|---|
| ||||
wget http://web.corral.tacc.utexas.edu/BioITeamBioinformaticsResource/CoreNGS/yeast_stuff/Sample_Yeast_L005_R1.cat.fastq.gz |
Now press Enter to get the command going. Repeat for the 2nd link. Check that you now see the two files (ls).
Copy from a corral location - cp or rsync
Suppose you have a corral allocation where your organization keeps its data, and that the sequencing data has been downloaded there. You can use various Linux commands to copy the data locally from there to your $SCRATCH area.
cp
The cp command copies one or more files from a local source to a local destination. It has the most common form:
cp [options] <source file 1> <source file 2> ... <destination directory>/
Make a directory in your scratch area and copy a single file to it. The trailing slash ( / ) on the destination says it is a directory.
| Tip |
|---|
By default wget creates a file in the current directory matching the last component of the URL (e.g. Sample_Yeast_L005_R1.cat.fastq.gz here). You can change the copied file name with wget's -O option. Also note that if you execute the same wget more than once, subsequent local files will be named with a .1, .2, etc. suffix. |
Copy from a corral location - cp or rsync
Suppose you have a corral allocation or stockyard area where your organization keeps its data, and that the sequencing data has been downloaded there. You can use various Linux commands to copy the data locally from there to your $SCRATCH area.
cp
The cp command copies one or more files from a local source to a local destination. It has the most common form:
cp [options] <source file 1> <source file 2> ... <destination directory>/
Make a directory in your Scratcharea and copy a single file to it. The trailing slash ( / ) on the destination says the destination is a directory.
| Code Block | ||||
|---|---|---|---|---|
| ||||
mkdir -p $SCRATCH | ||||
| Code Block | ||||
| ||||
mkdir -p $SCRATCH/data/test1 cp $CORENGS/misc/small.fq $SCRATCH/data/test1/ ls $SCRATCH/data/test1 # or.. mkdir -p ~/scratch/data/test1 cd ~/scratch/data/test1 cp $CORENGS/misc/small.fq . ls$SCRATCH/data/test1/ ls $SCRATCH/data/test1 # or.. mkdir -p ~/scratch/data/test1 cd ~/scratch/data/test1 cp $CORENGS/misc/small.fq . ls |
Copy an entire directory to your Scratch Copy an entire directory to your scratch area. The -r argument option says "recursive".
| Code Block | ||||
|---|---|---|---|---|
| ||||
mkdir -p $SCRATCH/data
cds
cd data
cp -r $CORENGS/general/ general/ |
...
rsync is a very complicated program, with many options (http://rsync.samba.org/ftp/rsync/rsync.html). However, if you use the recipe shown here for directories, it's hard to go wrong:
rsync -avP avW local/path/to/source_directory/ local/path/to/destination_directory/
Both the source and target directories are local (in some file system accessible directly from ls5 stampede2). Either full or relative path syntax can be used for both. The -avP avW options above stand for:
- -a means "archive mode", which implies the following options (and a few others)
- -p – preserve file permissions
- -t – preserve file times
- -l – copy symbolic links as links
- -r – recursively copy sub-directories
- -v means verbose
- -P means show ProgressW means transfer Whole file only
- Normally the rsync algorithm compares the contents of files that need to be copied and only transfers the different parts.
- For large files and binary files, figuring out what has changed (diff-ing) can take more time than just copying the whole file.
- The -W option disables file content comparisons (skips diff-ing).
Since these are all single-character options, they can be combined after one option prefix dash ( - ). Since these are all single-character options, they can be combined after one option prefix dash ( - ). You could also use options -ptlrvPptlrvW, separately, instead of using -a for "archive mode".
| Tip | ||
|---|---|---|
| ||
The trailing slash ( / ) on the source and destination directories are very important for rsync (and for other Linux copy commands also)! rsync will create the last directory level for you, but earlier levels must already exist. |
| Code Block | ||||
|---|---|---|---|---|
| ||||
| ||||
Another useful option is -W (copy Whole files only). In its normal mode, rsync will copy only changed parts of newer files. For large files, figuring out what has changed (diff-ing) can take more time than just copying the whole file. The rsync -W option says to skip diff-ing. | ||||
| ||||
mkdir -p $SCRATCH/data
cds
rsync -avW $CORENGS/ | ||||
| Code Block | ||||
| ||||
cds
rsync -avP $CORENGS/custom_tracks/ data/custom_tracks/ |
...
| Code Block | ||
|---|---|---|
| ||
rsync -avPavW /work/projects/BioITeam/projects/courses/Core_NGS_Tools/custom_tracks/ data/custom_tracks/ |
| Tip |
|---|
The bash shell has several convenient line editing features:
|
Copy from a remote computer - scp or rsync
...
Copy a single file to your $SCRATCH/data/test1 directory from the server named gapdhdragonfly.icmb.utexas.edu, using the user account corengstools. When prompted for a password, use the one we have written on to the board Zoom chat (or copy/paste the password from this file: $CORENGS/tacc/gapdhdragonfly_access.txt)
| Code Block | ||
|---|---|---|
| ||
cds cat $CORENGS/tacc/gapdhdragonfly_access.txt cds mkdir -p data/test2 scp corengstools@gapdhcorengstools@dragonfly.icmb.utexas.edu:~/custom_tracks/progeria_ctcf.vcf.gz ./data/test1test2/ ls ./data/test1test2 |
Notes:
- The 1st time you access a new host the SSH security prompt will appear
- You will be prompted for your remote host password
- The -r recursive argument works for scp also, just like for cp
...
- The tilde ( ~ ) at the start of the path means "relative to my home directory"We traverse through the BioITeam symbolic link created in your home directory earlier.
- We use the same tilde ( ~ ) in the destination to traverse the scratch symbolic link in your home directory.
...
- .
| Code Block | ||||
|---|---|---|---|---|
| ||||
rsync -ptlrvPavW corengstools@gapdhcorengstools@dragonfly.icmb.utexas.edu:~/custom_tracks/ ~/scratch/data/custom_tracks/ |
...
| Expand | ||
|---|---|---|
| ||
No, because all the source files were already present in the destination directory (you copied the same files earlier) with the same timestamps and file checksums. So rsync had nothing to do! |
...
| Code Block | ||
|---|---|---|
| ||
cd cp -r $CORENGS/work2/linuxpractice/what projects/BioITeam/projects/courses/Core_NGS_Tools/linuxpractice/what what # or using the $CORENGS environment variable cp -r $CORENGS/linuxpractice/what what cd what cat readme |
Where are you when you're all done?
| Expand | ||
|---|---|---|
| ||
|
step by step answers
| Expand | |||||
|---|---|---|---|---|---|
| |||||
From inside your ~/what directory:
|
| Expand | |||||
|---|---|---|---|---|---|
| |||||
From inside your ~/what/starts directory:
|
| Expand | |||||
|---|---|---|---|---|---|
| |||||
From inside your ~/what/starts/here directory:
|
| Expand | |||||
|---|---|---|---|---|---|
| |||||
From inside your ~/what/starts/here/changes directory:
|
...