$SCRATCH/core_ngs/references
| Table of Contents |
|---|
Overview
After raw sequence files are generated (in FASTQ format), quality-checked, and pre-processed in some way, the next step in most NGS pipelines is mapping to a reference genome.
...
Even though many mapping tools exist, a few individual programs have a dominant "market share" of the NGS world. In this section, we will primarily focus on two of the most versatile mappers: BWA and Bowtie2, the latter being part of the Tuxedo suite which includes the transcriptome-aware RNA-seq aligner Tophat2 as well as other downstream quantifiaction tools.
Connect to login5.ls5.tacc.utexas.edu
This should be second nature by now
Stage the alignment data
First stage the sample datasets and references we will use.
...
| File Name | Description | Sample |
|---|---|---|
| Sample_Yeast_L005_R1.cat.fastq.gz | Paired-end Illumina, First of pair, FASTQ | Yeast ChIP-seq |
| Sample_Yeast_L005_R2.cat.fastq.gz | Paired-end Illumina, Second of pair, FASTQ | Yeast ChIP-seq |
| human_rnaseq.fastq.gz | Paired-end Illumina, First of pair only, FASTQ | Human RNA-seq |
| human_mirnaseq.fastq.gz | Single-end Illumina, FASTQ | Human microRNA-seq |
| cholera_rnaseq.fastq.gz | Single-end Illumina, FASTQ | V. cholerae RNA-seq |
Reference Genomes
Before we get to alignment, we need a reference to align to. This is usually an organism's genome, but can also be any set of names sequences, such as a transcriptome or other set of genes.
...
| Tip | |||||
|---|---|---|---|---|---|
The BioITeam maintains a set of reference indexes for many common organisms and aligners. They can be found in aligner-specific sub-directories of the /work/projects/BioITeam/ref_genome area. E.g.:
|
Exercise #1: BWA global alignment – Yeast ChIP-seq
Overview ChIP-seq alignment workflow with BWA
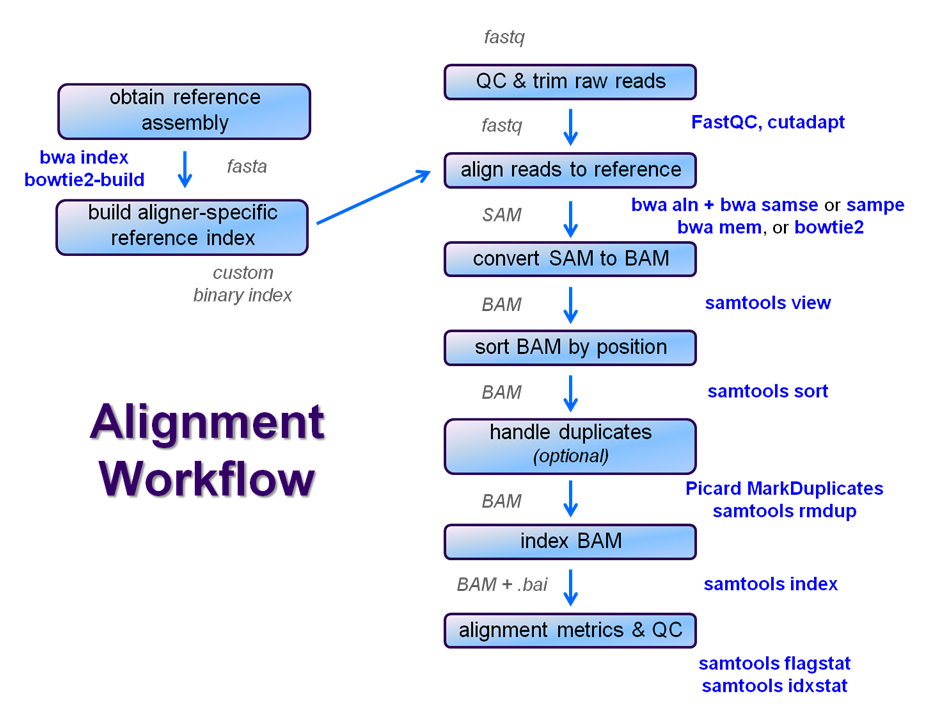
We will perform a global alignment of the paired-end Yeast ChIP-seq sequences using bwa. This workflow has the following steps:
...
We're going to skip the trimming step for now and see how it goes. We'll perform steps 2 - 5 now and leave samtools for the next exercise since steps 6 - 10 are common to nearly all post-alignment workflows.
Introducing BWA
Like other tools you've worked with so far, you first need to load bwa using the module system. Do that now, and then enter bwa with no arguments to view the top-level help page (many NGS tools will provide some help when called with no arguments).
...
As you can see, bwa include many sub-commands that perform the tasks we are interested in.
Building the BWA sacCer3 index
We will index the genome with the bwa index command. Type bwa index with no arguments to see usage for this sub-command.
...
| Code Block | ||
|---|---|---|
| ||
sacCer3.fa sacCer3.fa.amb sacCer3.fa.ann sacCer3.fa.bwt sacCer3.fa.pac sacCer3.fa.sa |
Exploring the FASTA with grep
It is often useful to know what chromosomes/contigs are in a FASTA file before you start an alignment so that you're familiar with the contig naming convention – and to verify that it's the one you expect. For example, chromosome 1 is specified differently in different references and organisms: chr1 (USCS human), chrI (UCSC yeast), or just 1 (Ensembl human GRCh37).
...
| Expand | ||
|---|---|---|
| ||
There are 17 contigs. |
Performing the bwa alignment
Now, we're ready to execute the actual alignment, with the goal of initially producing a SAM file from the input FASTQ files and reference. First prepare a directory for this exercise and link the sacCer3 reference directories there (this will make our commands more readable).
...
| Expand | ||
|---|---|---|
| ||
Both R1 and R2 reads must have separate alignment records, because there were 1,184,360 R1+R2 reads and the same number of alignment records. The SAM file must contain both mapped and un-mapped reads, because there were 1,184,360 R1+R2 reads and the same number of alignment records. Alignment records occur in the same read-name order as they did in the FASTQ, except that they come in pairs. The R1 read comes 1st, then the corresponding R2. This is called read name ordering. |
Using cut to isolate fields
Recall the format of a SAM alignment record:
...
| Expand | ||
|---|---|---|
| ||
| Recall that these are 100 bp reads and we did not remove adapter contamination. There will be a distribution of fragment sizes – some will be short – and those short fragments may not align without adapter removal (e.g. with fastx_trimmer). |
Exercise #2: Simple SAMtools Utilities
We have used several alignment methods that all generate results in the form of the near-universal SAM/BAM file format. The SAMtools program is a ubiquitously used set of tools that allow a user to manipulate SAM/BAM files in many different ways, ranging from simple tasks (like SAM/BAM interconversion) to more complex functions (like removal of PCR duplicates). It is available in the TACC module system in the typical fashion.
...
| Tip | ||
|---|---|---|
| ||
Be sure to check what version of samtools you are using! The most recent edition of SAMtools is 1.2, which has some important differences from the last version, 0.1.19. Most commands for this section are the same between the two versions, but if you see code from other sources using samtools, the version differences may be important. |
Samtools view
The utility samtools view provides a way of converting SAM (text format) files to BAM (binary, compressed) files directly. It also provides many, many other functions which we will discuss lster. To get a preview, execute samtools view without any other arguments. You should see:
...
| Code Block | ||
|---|---|---|
| ||
samtools view yeast_pairedend.bam | more |
Samtools sort
Look at the SAM file briefly using less. You will notice, if you scroll down, that the alignments are in no particular order, with chromosomes and start positions all mixed up. This makes searching through the file very inefficient. samtools sort is a piece of samtools that provides the ability to re-order entries in the SAM file either by coordinate position or by read name.
...
- The -O options says the output format should be BAM
- The -T options gives a prefix for temporary files produced during sorting
- By default sort writes its output to standard output, so we use > to redirect to a file named yeast_pairedend.sort.bam
Samtools index
Many tools (like the UCSC Genome Browser) only need to use sub-sections of the BAM file at a given point in time. For example, if you are viewing alignments that are within a particular gene alignments on other chromosomes generally do not need to be loaded. In order to speed up access, BAM files are indexed, producing BAI files which allow other programs to navigate directly to the alignments of interest. This is especially important when you have many alignments.
...
Most of the time when an index is required, it will be automatically located as long as it is in the same directory as its BAM file and shares the same name up until the .bai extension.
Samtools idxstats
Now that we have a sorted, indexed BAM file, we might like to get some simple statistics about the alignment run. For example, we might like to know how many reads aligned to each chromosome/contig. The samtools idxstats is a very simple tool that provides this information. If you type the command without any arguments, you will see that it could not be simpler - just type the following command:
...
| Tip |
|---|
If you're mapping to a non-genomic reference such as miRBase miRNAs or another set of genes (a transcriptome), samtools idxstats gives you a quick look at quantitative alignment results. |
Samtools flagstat
Finally, we might like to obtain some other statistics, such as the percent of all reads that aligned to the genome. The samtools flagstat tool provides very simple analysis of the SAM flag fields, which includes information like whether reads are properly paired, aligned or not, and a few other things. Its syntax is identical to that of samtools idxstats:
...
The most important statistic is the mapping rate, but this readout allows you to verify that some common expectations (e.g. that about the same number of R1 and R2 reads aligned, and that most mapped reads are proper pairs) are met.
Exercise #6: Yeast BWA PE alignment with Anna's script
Now that you've done everything the hard way, let's see how to do run an alignment pipeline using Anna's script.
...
| Code Block | ||
|---|---|---|
| ||
$path_code/script/align/align_bwa_illumina.sh ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz yeast_chip sacCer3 1 2>&1 | tee aln.yeast_chip.log |
Output files
This alignment pipeline script performs the following steps:
...
- aln.<prefix>.log – Log file of the entire alignment process.
- check the tail of this file to make sure the alignment was successful
- <prefix>.sort.dup.bam – Sorted, duplicate-marked alignment file.
- <prefix>.sort.dup.bam.bai – Index for the sorted, duplicate-marked alignment file
- <prefix>.samstats.txt – Summary alignment statistics from Anna's stats script
Verifying alignment success
The alignment log will have a "I ran successfully" message at the end if all went well, and if there was an error, the important information should also be at the end of the log file. So you can use tail to check the status of an alignment; for example:
...
The -L option tells grep to only print the filenames that don't contain the pattern. Perfect!
Checking alignment statistics
The <prefix>.samstats.txt statistics file produced by the alignment pipeline has a lot of good information in one place. If you use cat or more to view it you'll see this:
...
| Code Block |
|---|
delswr1_htz1_tap1t0.samstats.txt: Total mapped: 32761761 (86.8 %) delswr1_htz1_tap1t15.samstats.txt: Total mapped: 33699464 (89.2 %) delswr1_htz1_tap1t30.samstats.txt: Total mapped: 28441655 (87.6 %) wt_htz1_tap1t0.samstats.txt: Total mapped: 28454847 (89.5 %) wt_htz1_tap1t15.samstats.txt: Total mapped: 33245627 (90.9 %) wt_htz1_tap1t30.samstats.txt: Total mapped: 32567026 (90.7 %) |
TACC batch system considerations
The great thing about pipeline scripts like this is that you can perform alignments on many datasets in parallel at TACC.
...
| Expand | |||||
|---|---|---|---|---|---|
| |||||
Assuming you have $path_code set properly before submitting the job, the batch command would look like the command above, but you don't need the tee pipe. Instead, just redirect all output to a file. The example below shows how you would run alignments on two yeast samples in a batch file, adjusting the output prefix (yeast1, yeast2) and log file (aln.yeast1.log, aln.yeast2.log) accordingly.
|
x
Exercise #2: Bowtie2 global alignment - Vibrio cholerae RNA-seq
While we have focused on aligning eukaryotic data, the same tools can be used to perform identical functions with prokaryotic data. The major differences are less about the underlying data and much more about the external/public databases established to store and distribute reference data. For example, the Illumina iGenome resource provides pre-processed and uniform reference data, designed to be out-of-the-box compatible with aligners like bowtie2 and bwa. However, the limited number of available species are heavily biased towards model eukaryotes. If we wanted to study a prokaryote, the reference data must be downloaded from a resource like GenBank, and processed/indexed similarly to the procedure for mirbase.
While the alignment procedure for prokaryotes is broadly analogous, the reference preparation process is somewhat different, and will involve use of a biologically-oriented scripting library called BioPerl. In this exercise, we will use some RNA-seq data from Vibrio cholerae, published last year on GEO here, and align it to a reference genome.
Overview of Vibrio cholerae alignment workflow with Bowtie2
Alignment of this prokaryotic data follows the workflow below. Here we will concentrate on steps 1 and 2.
- Prepare the vibCho reference index for bowtie2 from a GenBank record using BioPerl
- Align reads using bowtie2, producing a SAM file
- Convert the SAM file to a BAM file (samtools view)
- Sort the BAM file by genomic location (samtools sort)
- Index the BAM file (samtools index)
- Gather simple alignment statistics (samtools flagstat and samtools idxstat)
Obtaining the GenBank record(s)
V. cholerae has two chromosomes. We download each separately.
- Navigate to http://www.ncbi.nlm.nih.gov/nuccore/NC_012582
- click on the Send down arrow (top right of page)
- select Complete Record
- select Clipboard as Destination
- click Add to Clipboard
- Perform these steps in your Terminal window
- mkdir $WORK/tmp; cd $WORK/tmp
- type wget then paste the URL from the clipboard
- Repeat steps 1 and 2 fot the 2nd chromosome
- NCBI URL is http://www.ncbi.nlm.nih.gov/nuccore/NC_012583
- you should now have 2 files, NC_012582 and NC_012583
- Combine the 2 files into one using cat
- cat NC_012582 NC_012583 > vibCho.gbk
Converting GenBank records into sequence (FASTA) and annotation (GFF) files
As noted earlier, many microbial genomes are available through repositories like GenBank that use specific file format conventions for storage and distribution of genome sequence and annotations. The GenBank file format is a text file that can be parsed to yield other files that are compatible with the pipelines we have been implementing.
...
After the header lines, each feature in the genome is represented by a line that gives chromosome, start, stop, strand, and other information. Features are things like "mRNA," "CDS," and "EXON." As you would expect in a prokaryotic genome it is frequently the case that the gene, mRNA, CDS, and exon annotations are identical, meaning they share coordinate information. You could parse these files further using commands like grep and awk to extract, say, all exons from the full file or to remove the header lines that begin with #.
Introducing bowtie2
Go ahead and load the bowtie2 module so we can examine some help pages and options. To do that, you must first load the perl module, and then the a specific version of bowtie2.
...
This is the key to using bowtie2 - it allows you to control almost everything about its behavior, but that also makes it is much more challenging to use than bwa – and it's easier to screw things up too!
Building the bowtie2 vibCho index
Before the alignment, of course, we've got to build a mirbase index using bowtie2-build (go ahead and check out its options). Unlike for the aligner itself, we only need to worry about a few things here:
...
This should also go pretty fast. You can see the resulting files using ls like before.
Performing the bowtie2 alignment
Now we will go back to our scratch area to do the alignment, and set up symbolic links to the index in the work area to simplify the alignment command:
...