...
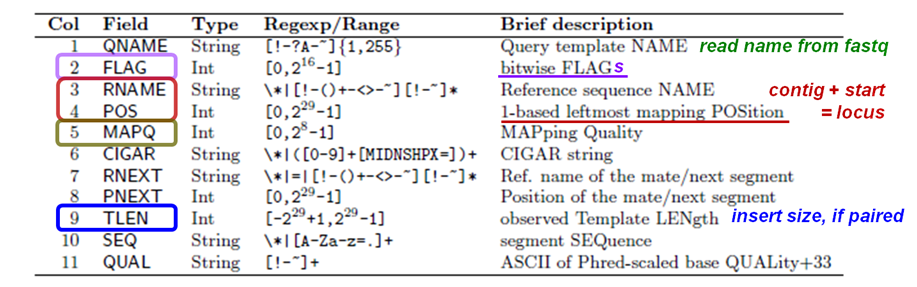
Recall the format of a SAM alignment record:
Suppose you wanted to look only at field 3 (contig name) values in the SAM file. You can do this with the handy cut command. Below is a simple example where you're asking cut to display the 3rd column value for the last 10 alignment records.
...
You can also specify a range of fields, and mix adjacent and non-adjacent fields. This displays fields 2 through 6, field 9, and all fields starting with the 12th (SAM tag fields).:
| Code Block | ||||
|---|---|---|---|---|
| ||||
tail -20 yeast_pairedend.sam | cut -f 2-6,9,12- |
You may have noticed that some alignment records contain contig names (e.g. chrV) in field 3 while others contain an asterisk ( * ). Usually the * means the record didn't map. (This isn't always true – later you'll see how to properly distinguish between mapped and unmapped reads using samtools.) We're going to use this heuristic along with cut to see about how many records represent aligned sequences.
...
| Expand | ||
|---|---|---|
| ||
| Recall that these are 100 bp reads and we did not remove adapter contamination. There will be a distribution of fragment sizes – some will be short – and those short fragments may not align without adapter removal (e.g. with fastx_trimmer). |
Exercise #2:
...
Basic SAMtools Utilities
We have used several alignment methods that all generate results in the form of the near-universal SAM/BAM file format. The The SAMtools program is a ubiquitously commonly used set of tools that allow a user to manipulate SAM/BAM files in many different ways, ranging from simple tasks (like SAM/BAM interconversion format conversion) to more complex functions (like removal of PCR duplicatessorting, indexing and statistics gathering). It is available in the TACC module system in the typical fashion.
In this exercise, we will use five very simple utilities provided by samtools: view, sort, index, flagstat, and idxstats. Each of these is executed in one line for a given SAM/BAM file. In the SAMtools/BEDtools section tomorrow we will explore samtools in more in depth.
For the sake of time and simplicity, here we are only going to run these commands on the yeast paired-end alignment file. The same commands can be run on the other files by changing the names, so feel free to try them on other SAM files. Indeed, it is very common in practice to use bash loops to generate many commands for a large set of alignments and deposit those commands into a batch job cmds file for submission.
To start, we will move to the directory containing our SAM files, among other things, and load up samtools using the module system. After loading it, just run the samtools command to see what the available tools are (and to see what the syntax of an actual command is).
| Code Block | ||
|---|---|---|
| ||
cd $SCRATCH/core_ngs/alignment
ls -la
module load samtools
samtools |
You will see the following screen after running samtools with no other options:
Load that module and see what samtools has to offer:
| Code Block | ||
|---|---|---|
| ||
module load samtools
samtools |
| Code Block | ||
|---|---|---|
| ||
Program: samtools (Tools for alignments in the SAM format)
Version: 1.3.1 (using htslib 1.3.1)
Usage: samtools <command> [options]
Commands:
-- Indexing
dict create a sequence dictionary file
faidx | ||
| Code Block | ||
Program: samtools (Tools for alignments in the SAM format) Version: 1.2 (using htslib 1.2.1) Usage: samtools <command> [options] Commands: -- indexing faidx index/extract FASTA index index index alignment -- editingEditing calmd calmd recalculate MD/NM tags and '=' bases fixmate fixmate fix mate information reheader reheader replace BAM header rmdup rmdup remove PCR duplicates targetcut targetcut cut fosmid regions (for fosmid pool only) addreplacerg adds or replaces RG tags -- fileFile operations collate bamshuf shuffle and group alignments by name cat cat concatenate BAMs merge merge merge sorted alignments mpileup mpileup multi-way pileup sort sort sort alignment file split split splits a file by read group quickcheck bam2fqquickly check if SAM/BAM/CRAM file appears intact fastq converts a BAM to a FASTQ fasta converts a BAM to a FASTA -- statsStatistics bedcov bedcov read depth per BED region depth depth compute the depth flagstat flagstat simple stats idxstats idxstats BAM index stats phase phase phase heterozygotes stats stats generate stats (former bamcheck) -- viewingViewing flags flags explain BAM flags tview tview text alignment viewer view view SAM<->BAM<->CRAM conversion |
| Tip | ||
|---|---|---|
| ||
Be sure to check what version of samtools you are using! The most recent edition of SAMtools is 1.2, which has some important differences from the last version, 0.1.19. Most commands for this section are the same between the two versions, but if you see code from other sources using samtools, the version differences may be important. |
Samtools view
depad convert padded BAM to unpadded BAM |
In this exercise, we will use five utilities provided by samtools: view, sort, index, flagstat, and idxstats. Each of these is executed in one line for a given SAM/BAM file. In the SAMtools/BEDtools sections tomorrow we will explore samtools in more in depth.
| Warning | ||
|---|---|---|
| ||
There are two main "eras" of SAMtools development:
Unfortunately, some functions with the same name in both version eras have different argument! So be sure you know which version you're using. (The samtools version is usually reported at the top of its usage listing). The default version in the ls5 module system is 1.3.1, but the BioITeam has a copy of the version 0.1.19 samtools for programs that might need it: /work/projects/BioITeam/ls5/bin/samtools-0.1.19. |
samtools view
The samtools view utility provides a way of converting between SAM (text) and BAM (binary, compressed) formatThe utility samtools view provides a way of converting SAM (text format) files to BAM (binary, compressed) files directly. It also provides many, many other functions which we will discuss lster. To get a preview, execute samtools view without any other arguments. You should see:
| Code Block | ||
|---|---|---|
| ||
Usage: samtools view [options] <in.bam>|<in.sam>|<in.cram> [region ...] Options: -b output BAM -C output CRAM (requires -T) -1 use fast BAM compression (implies -b) -u uncompressed BAM output (implies -b) -h include header in SAM output -H print SAM header only (no alignments) -c -c printprint only the count of matching records -o FILE output file name [stdout] -U FILE output reads not selected by filters to FILE [null] -t FILE FILE listing reference names and lengths (see long help) [null] -T FILE reference sequence FASTA FILE [null] -L FILE only include reads overlapping this BED FILE [null] -r STR only include reads in read group STR [null] -R FILE only include reads with read group listed in FILE [null] -q INT only include reads with mapping quality >= INT [0] -l STR only include reads in library STR [null] -m INT only include reads with number of CIGAR operations consuming consuming query sequence >= INT [0] -f INT only include reads with all bits set in INT set in FLAG [0] -F INT -F INT only only include reads with none of the bits set in INT set in FLAG [0] -x STR read tag to strip (repeatable) [null] -B collapse the backward CIGAR operation -s FLOAT integer part sets seed of random number generator [0]; rest sets fraction of templates to subsample [no subsampling] -@, --threads INT -@ INT number of BAM/CRAM compression threads [0] -? print long help, including note about region specification -S ignored (input format is auto-detected) |
That is a lot to process! For now, we just want to read in a SAM file and output a BAM file. The input format is auto-detected, so we don't need to say that we're inputing a SAM instead of a BAM. We just need to tell the tool to output the file in BAM format, and provide the name of the destination BAM file. This command is as follows:
| Code Block | ||
|---|---|---|
| ||
samtools view -b yeast_pairedend.sam -o yeast_pairedend.bam |
- the -b option tells the tool to output BAM format
- the -o option specifies the name of the output BAM file that will be created
How do you look at the BAM file contents now? That's simple. Just use samtools view. without the -b option. Remember to pipe output to a pager!
| Code Block | ||
|---|---|---|
| ||
samtools view yeast_pairedend.bam | more
|
Samtools sort
Look at the SAM file briefly using less. You will notice, if you scroll down, that the alignments are in no particular order, with chromosomes and start positions all mixed up. This makes searching through the file very inefficient. samtools sort is a piece of samtools that provides the ability to re-order entries in the SAM file either by coordinate position or by read name.
If you execute samtools sort without any options, you see its help page:
--input-fmt-option OPT[=VAL]
Specify a single input file format option in the form
of OPTION or OPTION=VALUE
-O, --output-fmt FORMAT[,OPT[=VAL]]...
Specify output format (SAM, BAM, CRAM)
--output-fmt-option OPT[=VAL]
Specify a single output file format option in the form
of OPTION or OPTION=VALUE
-T, --reference FILE
Reference sequence FASTA FILE [null] |
That is a lot to process! For now, we just want to read in a SAM file and output a BAM file. The input format is auto-detected, so we don't need to specify it (although you do in v0.1.19). We just need to tell the tool to output the file in BAM format.
| Code Block | ||
|---|---|---|
| ||
cd $SCRATCH/core_ngs/alignment/yeast_bwa
samtools view -b -o yeast_pairedend.bam yeast_pairedend.sam |
- the -b option tells the tool to output BAM format
- the -o option specifies the name of the output BAM file that will be created
How do you look at the BAM file contents now? That's simple. Just use samtools view without the -b option. Remember to pipe output to a pager!
| Code Block | ||
|---|---|---|
| ||
samtools view yeast_pairedend.bam | more
|
Notice that this does not show us the header record we saw at the start of the SAM file.
Exercise: What samtools view option will include the header records in its output? Which option would show only the header records?
| Expand | ||
|---|---|---|
| ||
samtools view -h shows header records along with alignment records. samtools view -H shows header records only. |
samtools sort
Looking at some of the alignment record information (e.g. samtools view yeast_pairedend.bam | cut -f 1-4 | more), you will notice that read names appear in adjacent pairs (for the R1 and R2), and the mappings are in no particular order, with chromosomes and start positions all mixed up. This makes searching through the file very inefficient. samtools sort provides the ability to re-order entries in the SAM file either by coordinate position or by read name.
If you execute samtools sort without any options, you see its help page:
| Code Block | ||
|---|---|---|
| ||
Usage: samtools sort [options...] [in.bam]
Options:
-l INT Set compression level, from 0 (uncompressed) to 9 (best)
-m INT Set maximum memory per thread; suffix K/M/G recognized [768M]
-n Sort by read name
-o FILE | ||
| Code Block | ||
Usage: samtools sort [options...] [in.bam] Options: -l INT Set compression level, from 0 (uncompressed) to 9 (best) -m INT Set maximum memory per thread; suffix K/M/G recognized [768M] -n Sort by read name -o FILE Write final output to FILE rather than standard output -O FORMAT Write output as FORMAT ('sam'/'bam'/'cram') (either -O or -T PREFIX Write temporary files to PREFIX.nnnn.bam -T is required) -@ INT Set number of sorting and compression threads [1] Legacy usage: samtools sort [options...] <in.bam> <out.prefix> Options: -f Use <out.prefix> as full final filename rather than prefix -o Write final output to stdoutFILE rather than <out.prefix>.bam standard output -l,m,n,@ Similar to corresponding options above |
In most cases you will be sorting a BAM file by position rather than by name. You can use either -o or reidrection with > to control the output.
To sort the paired-end yeast BAM file by coordinate, and get a BAM file named yeast_pairedend.sort.bam as output, execute the following command:
| Code Block | ||
|---|---|---|
| ||
samtools sort -O bam -T yeast_pairedend.sort yeast_pairedend.bam > yeast_pairedend.sort.bam |
- The -O options says the output format should be BAM
- The -T options gives a prefix for temporary files produced during sorting
- By default sort writes its output to standard output, so we use > to redirect to a file named yeast_pairedend.sort.bam
Samtools index
Many tools (like the UCSC Genome Browser) only need to use sub-sections of the BAM file at a given point in time. For example, if you are viewing alignments that are within a particular gene alignments on other chromosomes generally do not need to be loaded. In order to speed up access, BAM files are indexed, producing BAI files which allow other programs to navigate directly to the alignments of interest. This is especially important when you have many alignments.
The utility samtools index creates an index that has the exact name as the input BAM file, with suffix .bai appended. The help page, if you execute samtools index with no arguments, is as follows:
| Code Block |
|---|
Usage: samtools index [-bc] [-m INT] <in.bam> [out.index] Options: -bT PREFIX Write temporary files to PREFIX.nnnn.bam -@, --threads INT Set number of sorting and compression threads [1] --input-fmt-option OPT[=VAL] Specify a single input file format option in the form of OPTION or OPTION=VALUE -O, --output-fmt FORMAT[,OPT[=VAL]]... Specify output format (SAM, BAM, CRAM) --output-fmt-option OPT[=VAL] Generate BAI-format index for BAM files [default] Specify a -csingle output file format option in the form Generate CSI-format index for BAM files -m INT Set minimum interval size for CSI indices to 2^INT [14] |
Here, the syntax is way, way easier. We want a BAI-format index which is the default. (CSI-format is used with extremely long contigs, which we aren't considering here - the most common use case are highly polyploid plant genomes).
of OPTION or OPTION=VALUE
--reference FILE
Reference sequence FASTA FILE [null |
In most cases you will be sorting a BAM file from name order to coordinate order. You can use either -o or reidrection with > to control the output.
To sort the paired-end yeast BAM file by coordinate, and get a BAM file named yeast_pairedend.sort.bam as output, execute the following commandSo all we have to type is:
| Code Block | ||
|---|---|---|
| ||
samtools index sort -O bam -T yeast_pairedend.sort.bam |
...
tmp yeast_pairedend.bam > yeast_pairedend.sort.bam |
...
- The -O options says the output format should be BAM
- The -T options gives a prefix for temporary files produced during sorting
- sorting large BAMs will produce many temporary files during processing
- By default sort writes its output to standard output, so we use > to redirect to a file named yeast_pairedend.sort.bam
Samtools index
Many tools (like the UCSC Genome Browser) only need to use sub-sections of the BAM file at a given point in time. For example, if you are viewing alignments that are within a particular gene alignments on other chromosomes generally do not need to be loaded. In order to speed up access, BAM files are indexed, producing BAI files which allow other programs to navigate directly to the alignments of interest. This is especially important when you have many alignments.
The utility samtools index creates an index that has the exact name as the input BAM file, with suffix .bai appended. The help page, if you execute samtools index with no arguments, is as follows:
| Code Block |
|---|
Usage: samtools index [-bc] [-m INT] <in.bam> [out.index]
Options:
-b Generate BAI-format index for BAM files [default]
-c Generate CSI-format index for BAM files
-m INT Set minimum interval size for CSI indices to 2^INT [14] |
Here, the syntax is way, way easier. We want a BAI-format index which is the default. (CSI-format is used with extremely long contigs, which we aren't considering here - the most common use case are highly polyploid plant genomes).
So all we have to type is
...
Most of the time when an index is required, it will be automatically located as long as it is in the same directory as its BAM file and shares the same name up until the .bai extension.
Samtools idxstats
Now that we have a sorted, indexed BAM file, we might like to get some simple statistics about the alignment run. For example, we might like to know how many reads aligned to each chromosome/contig. The samtools idxstats is a very simple tool that provides this information. If you type the command without any arguments, you will see that it could not be simpler - just type the following command:
| Code Block | ||
|---|---|---|
| ||
samtools idxstats yeast_pairedend.sort.bam |
The output is a text file with four tab-delimited columns with the following meanings:
- chromosome name
- chromosome length
- number of mapped reads
- number of unmapped reads
The reason that the "unmapped reads" field for the named chromosomes is not zero is that, if one half of a pair of reads aligns while the other half does not, the unmapped read is still assigned to the chromosome its mate mapped to, but is flagged as unmapped.
| Tip |
|---|
If you're mapping to a non-genomic reference such as miRBase miRNAs or another set of genes (a transcriptome), samtools idxstats gives you a quick look at quantitative alignment results. |
Samtools flagstat
Finally, we might like to obtain some other statistics, such as the percent of all reads that aligned to the genome. The samtools flagstat tool provides very simple analysis of the SAM flag fields, which includes information like whether reads are properly paired, aligned or not, and a few other things. Its syntax is identical to that of samtools idxstats:
| Code Block | ||
|---|---|---|
| ||
samtools flagstatindex yeast_pairedend.sort.bam |
You should see something like this:
| Code Block |
|---|
1184360 + 0 in total (QC-passed reads + QC-failed reads)
0 + 0 secondary
0 + 0 supplementary
0 + 0 duplicates
547664 + 0 mapped (46.24%:-nan%)
1184360 + 0 paired in sequencing
592180 + 0 read1
592180 + 0 read2
473114 + 0 properly paired (39.95%:-nan%)
482360 + 0 with itself and mate mapped
65304 + 0 singletons (5.51%:-nan%)
534 + 0 with mate mapped to a different chr
227 + 0 with mate mapped to a different chr (mapQ>=5) |
Ignore the "+ 0" addition to each line - that is a carry-over convention for counting QA-failed reads that is no longer necessary.
The most important statistic is the mapping rate, but this readout allows you to verify that some common expectations (e.g. that about the same number of R1 and R2 reads aligned, and that most mapped reads are proper pairs) are met.
Exercise #6: Yeast BWA PE alignment with Anna's script
Now that you've done everything the hard way, let's see how to do run an alignment pipeline using Anna's script.
First the setup:
| Code Block | ||
|---|---|---|
| ||
mkdir -p $SCRATCH/core_ngs/align2/fastq
cd $SCRATCH/core_ngs/align2/fastq
cp /corral-repl/utexas/BioITeam/core_ngs_tools/alignment/*fastq.gz . |
Before executing the script you need to have one environment variable set. We'll do it at the command line here, but you could put it in your .bashrc file.
| Code Block | ||
|---|---|---|
| ||
export path_code=/work/01063/abattenh/code |
Now change into the directory and call the script with no arguments to see usage
| Code Block | ||
|---|---|---|
| ||
cd $SCRATCH/core_ngs/align2
$path_code/script/align/align_bwa_illumina.sh |
There are lots of bells and whistles in the arguments, but the most important are the first few:
- FASTQ file – full or relative path to the FASTQ file (just the R1 fastq if paired end). Can be compressed (.gz)
- output prefix – prefix for all the output files produced by the script. Should relate back to what the data is.
- assembly – genome assembly to use.
- there are pre-built indexes for some common eukaryotes (hg19, hg18, mm10, mm9, danRer7, sacCer3) that you can use
- or provide a full path for a bwa reference index you have built somewhere
- paired flag – 0 means single end (the default); 1 means paired end
- hard trim length – if you want the FASTQ hard trimmed down to a specific length, supply that number here
Now run the pipeline. By piping the output to tee <filename> we can see the script's progress at the terminal, and it also is written to <filename>.
| Code Block | ||
|---|---|---|
| ||
$path_code/script/align/align_bwa_illumina.sh ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz yeast_chip sacCer3 1 2>&1 | tee aln.yeast_chip.log |
Output files
This alignment pipeline script performs the following steps:
- Hard trims FASTQ, if optionally specified (fastx_trimmer)
- Aligns the R1 FASTQ (bwa aln)
- Aligns the R2 FASTQ, if paired end alignment specified (bwa aln)
- Reports the alignments as SAM (bwa samse for single end, or bwa sampe for paired end)
- Converts SAM to BAM (samtools view)
- Sorts the BAM (samtools sort)
- Marks duplicates (Picard MarkDuplicates)
- Indexes the sorted, duplicate-marked BAM (samtools index)
- Gathers statistics (samtools idxstats, samtools flagstat, plus a custom statistics script of Anna's)
- Removes intermediate files
There are a number of output files, with the most important being those desribed below.
- aln.<prefix>.log – Log file of the entire alignment process.
- check the tail of this file to make sure the alignment was successful
- <prefix>.sort.dup.bam – Sorted, duplicate-marked alignment file.
- <prefix>.sort.dup.bam.bai – Index for the sorted, duplicate-marked alignment file
- <prefix>.samstats.txt – Summary alignment statistics from Anna's stats script
Verifying alignment success
The alignment log will have a "I ran successfully" message at the end if all went well, and if there was an error, the important information should also be at the end of the log file. So you can use tail to check the status of an alignment; for example:
| Code Block | ||||
|---|---|---|---|---|
| ||||
tail aln.yeast_chip.log |
This will show something like:
This will produce a file named yeast_pairedend.bam.bai.
Most of the time when an index is required, it will be automatically located as long as it is in the same directory as its BAM file and shares the same name up until the .bai extension.
Samtools idxstats
Now that we have a sorted, indexed BAM file, we might like to get some simple statistics about the alignment run. For example, we might like to know how many reads aligned to each chromosome/contig. The samtools idxstats is a very simple tool that provides this information. If you type the command without any arguments, you will see that it could not be simpler - just type the following command:
| Code Block | ||
|---|---|---|
| ||
samtools idxstats yeast_pairedend.sort.bam |
The output is a text file with four tab-delimited columns with the following meanings:
- chromosome name
- chromosome length
- number of mapped reads
- number of unmapped reads
The reason that the "unmapped reads" field for the named chromosomes is not zero is that, if one half of a pair of reads aligns while the other half does not, the unmapped read is still assigned to the chromosome its mate mapped to, but is flagged as unmapped.
| Tip |
|---|
If you're mapping to a non-genomic reference such as miRBase miRNAs or another set of genes (a transcriptome), samtools idxstats gives you a quick look at quantitative alignment results. |
Samtools flagstat
Finally, we might like to obtain some other statistics, such as the percent of all reads that aligned to the genome. The samtools flagstat tool provides very simple analysis of the SAM flag fields, which includes information like whether reads are properly paired, aligned or not, and a few other things. Its syntax is identical to that of samtools idxstats:
| Code Block | ||
|---|---|---|
| ||
samtools flagstat yeast_pairedend.sort.bam |
You should see something like this:
| Code Block |
|---|
1184360 + 0 in total (QC-passed reads + QC-failed reads)
0 + 0 secondary
0 + 0 supplementary
0 + 0 duplicates
547664 + 0 mapped (46.24%:-nan%)
1184360 + 0 paired in sequencing
592180 + 0 read1
592180 + 0 read2
473114 + 0 properly paired (39.95%:-nan%)
482360 + 0 with itself and mate mapped
65304 + 0 singletons (5.51%:-nan%)
534 + 0 with mate mapped to a different chr
227 + 0 with mate mapped to a different chr (mapQ>=5) |
Ignore the "+ 0" addition to each line - that is a carry-over convention for counting QA-failed reads that is no longer necessary.
The most important statistic is the mapping rate, but this readout allows you to verify that some common expectations (e.g. that about the same number of R1 and R2 reads aligned, and that most mapped reads are proper pairs) are met.
Exercise #3: Yeast BWA PE alignment with BioITeam alignment script
Now that you've done everything the hard way, let's see how to do run an alignment pipeline using a BWA alignment script.
First the setup:
| Code Block | ||
|---|---|---|
| ||
mkdir -p $SCRATCH/core_ngs/align2/fastq
cd $SCRATCH/core_ngs/align2/fastq
cp /corral-repl/utexas/BioITeam/core_ngs_tools/alignment/*fastq.gz . |
Before executing the script you need to have one environment variable set. We'll do it at the command line here, but you could put it in your .bashrc file.
| Code Block | ||
|---|---|---|
| ||
export path_code=/work/01063/abattenh/code |
Now change into the directory and call the script with no arguments to see usage
| Code Block | ||
|---|---|---|
| ||
cd $SCRATCH/core_ngs/align2
$path_code/script/align/align_bwa_illumina.sh |
There are lots of bells and whistles in the arguments, but the most important are the first few:
- FASTQ file – full or relative path to the FASTQ file (just the R1 fastq if paired end). Can be compressed (.gz)
- output prefix – prefix for all the output files produced by the script. Should relate back to what the data is.
- assembly – genome assembly to use.
- there are pre-built indexes for some common eukaryotes (hg19, hg18, mm10, mm9, danRer7, sacCer3) that you can use
- or provide a full path for a bwa reference index you have built somewhere
- paired flag – 0 means single end (the default); 1 means paired end
- hard trim length – if you want the FASTQ hard trimmed down to a specific length, supply that number here
Now run the pipeline. By piping the output to tee <filename> we can see the script's progress at the terminal, and it also is written to <filename>.
| Code Block | ||
|---|---|---|
| ||
$path_code/script/align/align_bwa_illumina.sh ./fastq/Sample_Yeast_L005_R1.cat.fastq.gz yeast_chip sacCer3 1 2>&1 | tee aln.yeast_chip.log |
Output files
This alignment pipeline script performs the following steps:
- Hard trims FASTQ, if optionally specified (fastx_trimmer)
- Aligns the R1 FASTQ (bwa aln)
- Aligns the R2 FASTQ, if paired end alignment specified (bwa aln)
- Reports the alignments as SAM (bwa samse for single end, or bwa sampe for paired end)
- Converts SAM to BAM (samtools view)
- Sorts the BAM (samtools sort)
- Marks duplicates (Picard MarkDuplicates)
- Indexes the sorted, duplicate-marked BAM (samtools index)
- Gathers statistics (samtools idxstats, samtools flagstat, plus a custom statistics script of Anna's)
- Removes intermediate files
There are a number of output files, with the most important being those desribed below.
- aln.<prefix>.log – Log file of the entire alignment process.
- check the tail of this file to make sure the alignment was successful
- <prefix>.sort.dup.bam – Sorted, duplicate-marked alignment file.
- <prefix>.sort.dup.bam.bai – Index for the sorted, duplicate-marked alignment file
- <prefix>.samstats.txt – Summary alignment statistics from Anna's stats script
Verifying alignment success
The alignment log will have a "I ran successfully" message at the end if all went well, and if there was an error, the important information should also be at the end of the log file. So you can use tail to check the status of an alignment; for example:
| Code Block | ||||
|---|---|---|---|---|
| ||||
tail aln.yeast_chip.log |
This will show something like:
| Code Block |
|---|
..samstats file 'yeast_chip.samstats.txt' exists Thu May 28 16:36:01 CDT 2015
..samstats file file 'yeast_chip.samstats.txt' size ok Thu May 28 16:36:01 CDT 2015
--- |
| Code Block |
..samstats file 'yeast_chip.samstats.txt' exists Thu May 28 16:36:01 CDT 2015 ..samstats file file 'yeast_chip.samstats.txt' size ok Thu May 28 16:36:01 CDT 2015 --------------------------------------------------------- Cleaning up files... --------------------------------------------------------- ckRes 0 cleanup --------------------------------------------------------- AllCleaning bwa alignment tasks completed successfully! Thu May 28 16:36:01 CDT 2015 up files... --------------------------------------------------------- ckRes 0 cleanup -------------------- |
Notice that success message: "All bwa alignment tasks completed successfully!". It should only appear once in any successful alignment log.
When multiple alignment commands are run in parallel it is important to check them all, and you can use grep looking for part of the unique success message to do this.
For example, suppose I have run 6 alignments and have these 6 log files:
| Code Block |
|---|
aln.delswr1_htz1_tap1t0.log aln.delswr1_htz1_tap1t30.log aln.wt_htz1_tap1t15.log
aln.delswr1_htz1_tap1t15.log aln.wt_htz1_tap1t0.log aln.wt_htz1_tap1t30.log |
I can check that all 6 completed with this command:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep 'completed successfully' aln.*.log | wc -l |
If this command returns 6, I'm done. But what if it doesn't? If you grep -v (lines that don't contain the pattern), you'll get every line in every log file except the success message line, which is not what you want at all.
You could tail the log files one by one to see which one(s) don't have the message, but you can also use a special grep option to do this work:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep -L 'completed successfully' aln.*.log |
The -L option tells grep to only print the filenames that don't contain the pattern. Perfect!
Checking alignment statistics
The <prefix>.samstats.txt statistics file produced by the alignment pipeline has a lot of good information in one place. If you use cat or more to view it you'll see this:
-------------------------------------
All bwa alignment tasks completed successfully!
Thu May 28 16:36:01 CDT 2015
--------------------------------------------------------- |
Notice that success message: "All bwa alignment tasks completed successfully!". It should only appear once in any successful alignment log.
When multiple alignment commands are run in parallel it is important to check them all, and you can use grep looking for part of the unique success message to do this.
For example, suppose I have run 6 alignments and have these 6 log files:
| Code Block |
|---|
aln.delswr1_htz1_tap1t0.log aln.delswr1_htz1_tap1t30.log aln.wt_htz1_tap1t15.log
aln.delswr1_htz1_tap1t15.log aln.wt_htz1_tap1t0.log aln.wt_htz1_tap1t30.log |
I can check that all 6 completed with this command:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep 'completed successfully' aln.*.log | wc -l |
If this command returns 6, I'm done. But what if it doesn't? If you grep -v (lines that don't contain the pattern), you'll get every line in every log file except the success message line, which is not what you want at all.
You could tail the log files one by one to see which one(s) don't have the message, but you can also use a special grep option to do this work:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep -L 'completed successfully' aln.*.log |
The -L option tells grep to only print the filenames that don't contain the pattern. Perfect!
Checking alignment statistics
The <prefix>.samstats.txt statistics file produced by the alignment pipeline has a lot of good information in one place. If you use cat or more to view it you'll see this:
| Code Block |
|---|
-----------------------------------------------
Aligner: bwa
Total sequences: 1184360
Total mapped: 547664 (46.2 %)
Total unmapped: 636696 (53.8 %)
Primary: 547664 ( |
| Code Block |
----------------------------------------------- Aligner: bwa Total sequences: 1184360 Total mapped: 547664 (46.2 %) Total unmapped: 636696 (53.8 %) Primary: 547664 (100.0 %) Secondary: Duplicates: 324280 (59.2 %) Fwd strand: 272898 (49.8 %) Rev strand: 274766 (50.2 %) Multi hit: 18688 (3.4 %) Soft clip: 222451 (40.6 %) All match: 319429 (58.3 %) Indels: 6697 (1.2 %) Spliced: ----------------------------------------------- Total PE seqs: 1184360 PE seqs mapped: 547664 (46.2 %) Num PE pairs: 592180 F5 1st end mapped: 300477 (50.7 %) F3 2nd end mapped: 247187 (41.7 %) PE pairs mapped: 241180 (40.7 %) PE proper pairs: 236557 (39.9 %) ----------------------------------------------- Insert size stats for: yeast_chip Number of pairs: 236557 (proper) Number of insert sizes: 212 Mean [-/+ 1 SD]: 215 [153 277] (sd 62) Mode [Fivenum]: 223 [105 210 220 229 321] ----------------------------------------------- |
Since this was a paired end alignment there is paired-end specific information reported, including insert size statistics: mean/standard deviation, mode (most common insert size value) and fivenum (min, q1, median, q3 max insert sizes).
A quick way to check alignment stats if you have run multiple alignments is again to use grep. For example, for the 6 alignment files shown earlier, running this:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep 'Total map' *samstats.txt |
will produce output like this:
| Code Block |
|---|
delswr1_htz1_tap1t0.samstats.txt: Total mapped: 32761761 (86.8 %)
delswr1_htz1_tap1t15.samstats.txt: Total mapped: 33699464 (89.2 %)
delswr1_htz1_tap1t30.samstats.txt: Total mapped: 28441655 (87.6 %)
wt_htz1_tap1t0.samstats.txt: Total mapped: 28454847 (89.5 %)
wt_htz1_tap1t15.samstats.txt: Total mapped: 33245627 (90.9 %)
wt_htz1_tap1t30.samstats.txt: Total mapped: 32567026 (90.7 %) |
TACC batch system considerations
The great thing about pipeline scripts like this is that you can perform alignments on many datasets in parallel at TACC.
Anna's alignment pipeline scripts are written to take advantage of having multiple cores on TACC nodes, and are thus designed to run with at most two pipeline commands per TACC node.
| Tip | ||
|---|---|---|
| ||
These pipeline scripts should always be run with a wayness of 2 (-w 2) in the TACC batch system, meaning two commands per node. |
Assuming you have your alignment commands in a file called aln.cmds, here's how to create and submit a batch job for the commands.
| Code Block | ||||
|---|---|---|---|---|
| ||||
launcher_creator.py -n aln -j aln.cmds -t 12:00:00 -q normal -w 2
sbatch aln.slurm
showq -u |
Note the maximum run time specified here is 12 hours (-t 12:00:00). This is a reasonable value for a higher eukaryote with 20-40 M reads, and is way more than a yeast alignment would need (~ 4 hours). For very deeply sequenced eukaryotes (e.g. human genome re-sequencing with hundresd of millions of reads), you may want to specify the maximum job time of 48 hours.
Exercise: What would alignment commands look like if you were putting it in a batch system .cmds file?
| Expand | |||||
|---|---|---|---|---|---|
| |||||
Assuming you have $path_code set properly before submitting the job, the batch command would look like the command above, but you don't need the tee pipe. Instead, just redirect all output to a file. The example below shows how you would run alignments on two yeast samples in a batch file, adjusting the output prefix (yeast1, yeast2) and log file (aln.yeast1.log, aln.yeast2.log) accordingly.
|
x
Exercise #2: Bowtie2 global alignment - Vibrio cholerae RNA-seq
While we have focused on aligning eukaryotic data, the same tools can be used to perform identical functions with prokaryotic data. The major differences are less about the underlying data and much more about the external/public databases established to store and distribute reference data. For example, the Illumina iGenome resource provides pre-processed and uniform reference data, designed to be out-of-the-box compatible with aligners like bowtie2 and bwa. However, the limited number of available species are heavily biased towards model eukaryotes. If we wanted to study a prokaryote, the reference data must be downloaded from a resource like GenBank, and processed/indexed similarly to the procedure for mirbase.
While the alignment procedure for prokaryotes is broadly analogous, the reference preparation process is somewhat different, and will involve use of a biologically-oriented scripting library called BioPerl. In this exercise, we will use some RNA-seq data from Vibrio cholerae, published last year on GEO here, and align it to a reference genome.
Overview of Vibrio cholerae alignment workflow with Bowtie2
Alignment of this prokaryotic data follows the workflow below. Here we will concentrate on steps 1 and 2.
- Prepare the vibCho reference index for bowtie2 from a GenBank record using BioPerl
- Align reads using bowtie2, producing a SAM file
- Convert the SAM file to a BAM file (samtools view)
- Sort the BAM file by genomic location (samtools sort)
- Index the BAM file (samtools index)
- Gather simple alignment statistics (samtools flagstat and samtools idxstat)
Obtaining the GenBank record(s)
V. cholerae has two chromosomes. We download each separately.
- Navigate to http://www.ncbi.nlm.nih.gov/nuccore/NC_012582
- click on the Send down arrow (top right of page)
- select Complete Record
- select Clipboard as Destination
- click Add to Clipboard
- Perform these steps in your Terminal window
- mkdir $WORK/tmp; cd $WORK/tmp
- type wget then paste the URL from the clipboard
- Repeat steps 1 and 2 fot the 2nd chromosome
- NCBI URL is http://www.ncbi.nlm.nih.gov/nuccore/NC_012583
- you should now have 2 files, NC_012582 and NC_012583
- Combine the 2 files into one using cat
- cat NC_012582 NC_012583 > vibCho.gbk
Converting GenBank records into sequence (FASTA) and annotation (GFF) files
As noted earlier, many microbial genomes are available through repositories like GenBank that use specific file format conventions for storage and distribution of genome sequence and annotations. The GenBank file format is a text file that can be parsed to yield other files that are compatible with the pipelines we have been implementing.
Go ahead and look at some of the contents of a GenBank file with the following commands (execute these one at a time):
| Code Block | ||
|---|---|---|
| ||
cd $WORK/core_ngs/references
less vibCho.O395.gbk # use q to quit less
grep -A 5 ORIGIN vibCho.O395.gbk |
As the less command shows, the file begins with a description of the organism and some source information, and the contains annotations for each bacterial gene. The grep command shows that, indeed, there is sequence information here (flagged by the word ORIGIN) that could be exported into a FASTA file. There are a couple ways of extracting the information we want, namely the reference genome and the gene annotation information, but a convenient one (that is available through the module system at TACC) is BioPerl.
We load BioPerl like we have loaded other modules, with the caveat that we must load regular Perl before loading BioPerl:
| Code Block | ||
|---|---|---|
| ||
module load perl
module load bioperl |
These commands make several scripts directly available to you. The one we will use is called bp_seqconvert.pl, and it is a BioPerl script used to inter-convert file formats like FASTA, GBK, and others. This script produces two output files:
- a FASTA format file for indexing and alignment
- a GFF file (standing for General Feature Format) contains information about all genes (or, more generally, features) in the genome
- remember, annotations such as GFFs must always match the reference you are using
To see how to use the script, just execute:
| Code Block | ||
|---|---|---|
| ||
bp_seqconvert.pl |
Clearly, there are many file formats that we can use this script to convert. In our case, we are moving from genbank to fasta, so the commands we would execute to produce and view the FASTA files would look like this:
| Code Block | ||
|---|---|---|
| ||
cd $WORK/core_ngs/references
bp_seqconvert.pl --from genbank --to fasta < vibCho.O395.gbk > vibCho.O395.fa
mv vibCho.O395.fa fasta/
grep ">" fasta/vibCho.O395.fa
less fasta/vibCho.O395.fa |
Now we have a reference sequence file that we can use with the bowtie2 reference builder, and ultimately align sequence data against.
Recall from when we viewed the GenBank file that there are genome annotations available as well that we would like to extract into GFF format. However, the bp_seqconvert.pl script is designed to be used to convert sequence formats, not annotation formats. Fortunately, there is another script called bp_genbank2gff3.pl that can take a GenBank file and produce a GFF3 (the most recent format convention for GFF files) file. To run it and see the output, run these commands:
| Code Block | ||
|---|---|---|
| ||
bp_genbank2gff3.pl --format Genbank vibCho.O395.gbk
mv vibCho.O395.gbk.gff vibCho.O395.gff
less vibCho.O395.gff |
After the header lines, each feature in the genome is represented by a line that gives chromosome, start, stop, strand, and other information. Features are things like "mRNA," "CDS," and "EXON." As you would expect in a prokaryotic genome it is frequently the case that the gene, mRNA, CDS, and exon annotations are identical, meaning they share coordinate information. You could parse these files further using commands like grep and awk to extract, say, all exons from the full file or to remove the header lines that begin with #.
Introducing bowtie2
Go ahead and load the bowtie2 module so we can examine some help pages and options. To do that, you must first load the perl module, and then the a specific version of bowtie2.
| Code Block | ||
|---|---|---|
| ||
module load perl
module load bowtie/2.2.0
|
Now that it's loaded, check out the options. There are a lot of them! In fact for the full range of options and their meaning, Google "Bowtie2 manual" and bring up that page. The Table of Contents is several pages long! Ouch!
This is the key to using bowtie2 - it allows you to control almost everything about its behavior, but that also makes it is much more challenging to use than bwa – and it's easier to screw things up too!
Building the bowtie2 vibCho index
Before the alignment, of course, we've got to build a mirbase index using bowtie2-build (go ahead and check out its options). Unlike for the aligner itself, we only need to worry about a few things here:
- reference_in file is just the FASTA file containing mirbase v20 sequences
- bt2_index_base is the prefix of where we want the files to go
To build the reference index for alignment, we actually only need the FASTA file, since annotations are often not necessary for alignment. (This is not always true - extensively spliced transcriptomes requires splice junction annotations to align RNA-seq data properly, but for now we will only use the FASTA file.)
| Code Block | ||
|---|---|---|
| ||
mkdir -p $WORK/core_ngs/references/bt2/vibCho
mv $WORK/core_ngs/references/vibCho.O395.fa $WORK/core_ngs/references/fasta
cd $WORK/core_ngs/references/bt2/vibCho
ln -s -f ../../fasta/vibCho.O395.fa
ls -la |
Now build the index using bowtie2-build:
| Code Block | ||||
|---|---|---|---|---|
| ||||
bowtie2-build vibCho.O395.fa vibCho.O395 |
This should also go pretty fast. You can see the resulting files using ls like before.
Performing the bowtie2 alignment
Now we will go back to our scratch area to do the alignment, and set up symbolic links to the index in the work area to simplify the alignment command:
| Code Block | ||
|---|---|---|
| ||
cd $SCRATCH/core_ngs/alignment
ln -s -f $WORK/core_ngs/references/bt2/vibCho vibCho |
Note that here the data is from standard mRNA sequencing, meaning that the DNA fragments are typically longer than the reads. There is likely to be very little contamination that would require using a local rather than global alignment, or many other pre-processing steps (e.g. adapter trimming). Thus, we will run bowtie2 with default parameters, omitting options other than the input, output, and reference index.
As you can tell from looking at the bowtie2 help message, the general syntax looks like this:
| Code Block |
|---|
bowtie2 [options]* -x <bt2-idx> {-1 <m1> -2 <m2> | -U <r>} [-S <sam>] |
So our command would look like this:
| Code Block | ||
|---|---|---|
| ||
bowtie2 -x vibCho/vibCho.O395 -U fastq/cholera_rnaseq.fastq.gz -S cholera_rnaseq.sam |
| Expand | ||
|---|---|---|
| ||
Parameters are:
|
Create a commands file called bt2_vibCho.cmds with this task definition then generate and submit a batch job for it (time 1 hour, development queue).
| Expand | |||||||
|---|---|---|---|---|---|---|---|
| |||||||
Use nano to create the bt2_vibCho.cmds file. Then:
|
...
sizes: 212
Mean [-/+ 1 SD]: 215 [153 277] (sd 62)
Mode [Fivenum]: 223 [105 210 220 229 321]
----------------------------------------------- |
Since this was a paired end alignment there is paired-end specific information reported, including insert size statistics: mean/standard deviation, mode (most common insert size value) and fivenum (min, q1, median, q3 max insert sizes).
A quick way to check alignment stats if you have run multiple alignments is again to use grep. For example, for the 6 alignment files shown earlier, running this:
| Code Block | ||||
|---|---|---|---|---|
| ||||
grep 'Total map' *samstats.txt |
will produce output like this:
| Code Block |
|---|
delswr1_htz1_tap1t0.samstats.txt: Total mapped: 32761761 (86.8 %)
delswr1_htz1_tap1t15.samstats.txt: Total mapped: 33699464 (89.2 %)
delswr1_htz1_tap1t30.samstats.txt: Total mapped: 28441655 (87.6 %)
wt_htz1_tap1t0.samstats.txt: Total mapped: 28454847 (89.5 %)
wt_htz1_tap1t15.samstats.txt: Total mapped: 33245627 (90.9 %)
wt_htz1_tap1t30.samstats.txt: Total mapped: 32567026 (90.7 %) |
TACC batch system considerations
The great thing about pipeline scripts like this is that you can perform alignments on many datasets in parallel at TACC.
Anna's alignment pipeline scripts are written to take advantage of having multiple cores on TACC nodes, and are thus designed to run with at most two pipeline commands per TACC node.
| Tip | ||
|---|---|---|
| ||
These pipeline scripts should always be run with a wayness of 2 (-w 2) in the TACC batch system, meaning two commands per node. |
Assuming you have your alignment commands in a file called aln.cmds, here's how to create and submit a batch job for the commands.
| Code Block | ||||
|---|---|---|---|---|
| ||||
launcher_creator.py -n aln -j aln.cmds -t 12:00:00 -q normal -w 2
sbatch aln.slurm
showq -u |
Note the maximum run time specified here is 12 hours (-t 12:00:00). This is a reasonable value for a higher eukaryote with 20-40 M reads, and is way more than a yeast alignment would need (~ 4 hours). For very deeply sequenced eukaryotes (e.g. human genome re-sequencing with hundresd of millions of reads), you may want to specify the maximum job time of 48 hours.
Exercise: What would alignment commands look like if you were putting it in a batch system .cmds file?
| Expand | |||||
|---|---|---|---|---|---|
| |||||
Assuming you have $path_code set properly before submitting the job, the batch command would look like the command above, but you don't need the tee pipe. Instead, just redirect all output to a file. The example below shows how you would run alignments on two yeast samples in a batch file, adjusting the output prefix (yeast1, yeast2) and log file (aln.yeast1.log, aln.yeast2.log) accordingly.
|