Reservations
Use our summer school reservation (CoreNGS-Tue) when submitting batch jobs to get higher priority on the ls6 normal queue today:
sbatch --reservation=CoreNGS-Tue <batch_file>.slurm
idev -m 180 -N 1 -A OTH21164 -r CoreNGS-Tue
Note that the reservation name (CoreNGS-Tue) is different from the TACC allocation/project for this class, which is OTH21164.
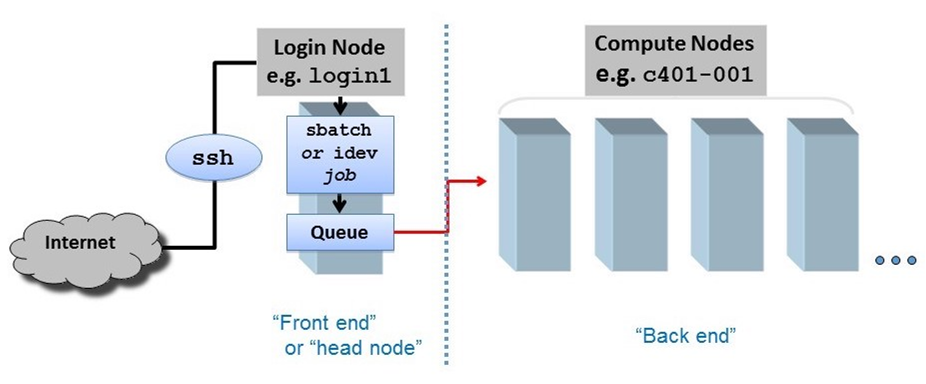
Compute cluster overview
When you SSH into ls6, your session is assigned to one of a small set of login nodes (also called head nodes). These are separate from the cluster compute nodes that will run your jobs.
Think of a node as a computer, like your laptop, but probably with more cores and memory. Now multiply that computer a thousand or more, and you have a cluster.
The small set of login nodes are a shared resource (type the users command to see everyone currently logged in) and are not meant for running interactive programs – for that you submit a description of what you want done to a batch system, which distributes the work to one or more compute nodes.
On the other hand, the login nodes are intended for copying files to and from TACC, so they have a lot of network bandwidth while compute nodes have limited network bandwidth.
So follow these guidelines:
- Do not perform substantial computation on the login nodes.
- They are closely monitored, and you will get warnings from the TACC admin folks!
- Code is usually developed and tested somewhere other than TACC, and only moved over when pretty solid.
- Do not perform significant network access from your batch jobs.
- Instead, stage your data from a login node onto $SCRATCH before submitting your job.
Lonestar6 and Stampede2 overview and comparison
Here is a comparison of the configurations and ls6 and stampede2. As you can see, stampede2 is the larger cluster, launched in 2017, but ls6, launched om 2022, has fewer but more powerful nodes.
| ls6 | stampede2 | |
|---|---|---|
| login nodes | 3 128 cores each | 6 28 cores each |
| standard compute nodes | 560 AMD Epyc Milan processors 128 cores per node | 4,200 KNL (Knights Landing) processors
1,736 SKX (Skylake) processors
|
| GPU nodes | 16 AMD Epyc Milan processors 128 cores per nod 2x NVIDIA A100 GPUs | -- |
| batch system | SLURM | SLURM |
| maximum job run time | 48 hours, normal queue 2 hours, development queue | 96 hours on KNL nodes, normal queue 48 hours on SKX nodes, normal queue 2 hours, development queue |
User guides for ls6 and stampede2 can be found at:
- https://portal.tacc.utexas.edu/user-guides/lonestar6
- https://portal.tacc.utexas.edu/user-guides/stampede2
Unfortunately, the TACC user guides are aimed towards a different user community – the weather modelers and aerodynamic flow simulators who need very fast matrix manipulation and other High Performance Computing (HPC) features. The usage patterns for bioinformatics – generally running 3rd party tools on many different datasets – is rather a special case for HPC. TACC calls our type of processing "parameter sweep jobs" and has a special process for running them, using their launcher module.
About cores and hyperthreads
Note the use of the term virtual core on stampede2. Compute cores are standalone processors – mini CPUs, each of which can execute separate sets of instructions. However modern cores may also have hyperthreading enabled, where a single core can appear as more than one virtual processor to the operating system (see https://en.wikipedia.org/wiki/Hyper-threading). For example, stampede2 nodes have 2 or 4 hyperthreads (HTs) per core. So KNL nodes with 4 HTs for each of the 68 physical cores, have a total of 272 virtual cores.
Threading is an operating system scheduling mechanism for allowing one CPU/core to execute multiple computations, seemingly in parallel.
The writer of a program that takes advantage of threading first identifies portions of code that can run in parallel because the computations are independent. The programmer assigns some number of threads to that work (usually based on a command-line option) using specific thread and synchronization programming language constructs. An example is the the samtools sort -@ N option to specify N threads can be used for sorting independent sets of the input alignments.
If there are multiple cores/CPUs available, the operating system can assign a program thread to each of them for actual parallelism. But only "seeming" (or virtual) parallelism occurs if there are fewer cores than the number of threads specified.
Suppose there's only one core/CPU. The OS assigns program thread A to the core to run until the program performs an I/O operation that causes it to be "suspended" for the I/O operation to complete. During this time, when normally the CPU would be doing nothing but waiting on the I/O to complete, the OS assigns program thread B to the CPU and lets it do some work. This threading allows more efficient use of existing cores as long as the multiple program threads being assigned do some amount of I/O or other operations that cause them to suspend. But trying to run multiple compute-only, no-I/O programs using multiple threads on one CPU just causes "thread thrashing" -- OS scheduler overhead when threads are suspended for time, not just I/O.
The analogy is a grocery store where there are 5 customers (threads). If there are 5 checkout lines (cores), each customer (thread) can be serviced in a separate checkout line (core). But if there's only one checkout line (core) open, the customers (threads) will have to wait in line. To be a more accurate analogy, any checkout clerk would be able to handle some part of checkout for each customer, then while waiting for the customer to find and enter credit card information, the clerk could handle a part of a different customer's checkout.
Hyperthreading is just a hardware implementation of OS scheduling. Each CPU offers some number of "virtual cores" (hyperthreads) that can "almost" act like separate cores using various hardware tricks. Still, if the work assigned to multiple hyperthreads on a single core does not pause from time to time, thread thrashing will occur.
Software at TACC
Programs and your $PATH
When you type in the name of an arbitrary program (ls for example), how does the shell know where to find that program? The answer is your $PATH. $PATH is a predefined environment variable whose value is a list of directories.The shell looks for program names in that list, in the order the directories appear.
To determine where the shell will find a particular program, use the which command. Note that which tells you where it looked if it cannot find the program.
which rsync which cat which bwa # not yet available to you
The module system
The module system is an incredibly powerful way to have literally thousands of software packages available, some of which are incompatible with each other, without causing complete havoc. The TACC staff stages packages in well-known locations that are NOT on your $PATH. Then, when a module is loaded, its binaries are added to your $PATH.
For example, the following module load command makes the singularity container management system available to you:
# first type "singularity" to show that it is not present in your environment: singularity # it's not on your $PATH either: which singularity # now add biocontainers to your environment and try again: module load biocontainers # and see how singularity is now on your $PATH: which singularity # you can see the new directory at the front of $PATH echo $PATH # to remove it, use "unload" module unload biocontainers singularity # gone from $PATH again... which singularity
TACC BioContainers modules
It is quite a large systems administration task to install software at TACC and configure it for the module system. As a result, TACC was always behind in making important bioinformatics software available. To address this problem, TACC moved to providing bioinformatics software via containers, which are virtual machines like VMware and Virtual Box, but are lighter weight: they require less disk space because they rely more on the host's base Linux environment. Specifically, TACC (and many other High Performance Computing clusters) use Singularity containers, which are similar to Docker containers but are more suited to the HPC environment (in fact one can build a Docker container then easily convert it to Singularity for use at TACC).
TACC obtains its containers from BioContainers (https://biocontainers.pro/ and https://github.com/BioContainers/containers), a large public repository of bioinformatics tool Singularity containers. This has allowed TACC to easily provision thousands of such tools!
These BioContainers are not visible in TACC's "standard" module system, but only after the master biocontainers module is loaded. Once it has been loaded, you can search for your favorite bioinformatics program using module spider.
# Verify that samtools is not available samtools # and cannot be found in the standard module system module spider samtools # Load the BioContainers master module (this takes a while) module load biocontainers # Now look for these programs module spider samtools module spider Rstats module spider kallisto module spider bowtie2 module spider minimap2 module spider multiqc module spider gatk module spider velvet
Notice how the BioContainers module names have "ctr" in their names, version numbers, and other identifying information.
The standard TACC module system has been phased out for bioinformatics programs, so always look for your application in BioContainers.
While it's great that there are now hundreds of programs available through BioContainers, the one drawback is that they can only be run on cluster compute nodes, not on login nodes. To test BioContainer program interactively, you will need to use TACC's idev command to obtain an interactive cluster node. More on this shortly...
loading a biocontainer module
Once the biocontainers module has been loaded, you can just module load the desired tool, as with the kallisto pseudo-aligner program below.
# Load the Biocontainers master module module load biocontainers # Verify kallisto is not yet available kallisto # Load the default kallisto biocontainer module load kallisto # Verify kallisto is not available (although not on login nodes) kallisto
Note that loading a BioContainer does not add anything to your $PATH. Instead, it defines an alias, which is just a shortcut for executing the command. You can see the alias definition using the type command. And you can ensure the program is available using the command -v utility.
# Note that kallisto has not been added to your $PATH, but instead has an alias which kallisto # Ensure kallisto is available with command -v command -v kallisto
installing custom software
Even with all the tools available at TACC, inevitably you'll need something they don't have. In this case you can build the tool yourself and install it in a local TACC directory. While building 3rd party tools is beyond the scope of this course, it's really not that hard. The trick is keeping it all organized.
For one thing, remember that your $HOME directory quota is fairly small (10 GB on ls6), and that can fill up quickly if you install many programs. We recommend creating an installation area in your $WORK directory and installing programs there. You can then make symbolic links to the binaries you need in your ~/local/bin directory (which was added to your $PATH in your .bashrc).
See how we used a similar trick to make the launcher_creator.py program available to you. Using the ls -l option shows you where symbolic links point to:
ls -l ~/local/bin # this will tell you the real location of the launcher_creator.py script is # /work/projects/BioITeam/common/bin/launcher_creator.py
$PATH caveat
Remember that the order of locations in the $PATH environment variable is the order in which the locations will be searched.
Job Execution
Job execution is controlled by the SLURM batch system on both stampede2 and ls6.
To run a job you prepare 2 files:
- a commands file file containing the commands to run, one task per line (<job_name>.cmds)
- a job control file that describes how to run the job (<job_name>.slurm)
The process of running the job involves these steps:
- Create a commands file containing exactly one task per line.
- Prepare a job control file for the commands file that describes how the job should be run.
- You submit the job control file to the batch system. The job is then said to be queued to run.
- The batch system prioritizes the job based on the number of compute nodes needed and the job run time requested.
- When compute nodes become available, the job tasks (command lines in the <job_name>.cmds file) are assigned to one or more compute nodes and begin to run in parallel.
- The job completes when either:
- you cancel the job manually
- all job tasks complete (successfully or not!)
- the requested job run time has expired
SLURM at a glance
Here are the main components of the SLURM batch system.
| stampede2, ls5 | |
|---|---|
| batch system | SLURM |
| batch control file name | <job_name>.slurm |
| job submission command | sbatch <job_name>.slurm |
| job monitoring command | showq -u |
| job stop command | scancel -n <job name> |
Simple example
Let's go through a simple example. Execute the following commands to copy a pre-made simple.cmds commands file:
mkdir -p $SCRATCH/core_ngs/slurm/simple cd $SCRATCH/core_ngs/slurm/simple cp $CORENGS/tacc/simple.cmds .
What are the tasks we want to do? Each task corresponds to one line in the simple.cmds commands file, so let's take a look at it using the cat (concatenate) command that simply reads a file and writes each line of content to standard output (here, your Terminal):
cat simple.cmds
The tasks we want to perform look like this:
sleep 5; echo "Command 1 on `hostname` - `date`" > cmd1.log 2>&1 sleep 5; echo "Command 2 on `hostname` - `date`" > cmd2.log 2>&1 sleep 5; echo "Command 3 on `hostname` - `date`" > cmd3.log 2>&1 sleep 5; echo "Command 4 on `hostname` - `date`" > cmd4.log 2>&1 sleep 5; echo "Command 5 on `hostname` - `date`" > cmd5.log 2>&1 sleep 5; echo "Command 6 on `hostname` - `date`" > cmd6.log 2>&1 sleep 5; echo "Command 7 on `hostname` - `date`" > cmd7.log 2>&1 sleep 5; echo "Command 8 on `hostname` - `date`" > cmd8.log 2>&1
There are 8 tasks. Each task sleeps for 5 seconds, then uses the echo command to output a string containing the task number and date to a log file named for the task number. Notice that we can put two commands on one line if they are separated by a semicolon ( ; ).
Use the handy launcher_creator.py program to create the job control file.
launcher_creator.py -j simple.cmds -n simple -t 00:01:00 -a OTH21164 -q development
You should see output something like the following, and you should see a simple.slurm batch submission file in the current directory.
Project simple. Using job file simple.cmds. Using development queue. For 00:01:00 time. Using OTH21164 allocation. Not sending start/stop email. Launcher successfully created. Type "sbatch simple.slurm" to queue your job.
Submit your batch job then check the batch queue to see the job's status.
sbatch simple.slurm
showq -u
# Output looks something like this:
-------------------------------------------------------------
Welcome to the Lonestar6 Supercomputer
-------------------------------------------------------------
--> Verifying valid submit host (login1)...OK
--> Verifying valid jobname...OK
--> Verifying valid ssh keys...OK
--> Verifying access to desired queue (normal)...OK
--> Checking available allocation (OTH21164)...OK
Submitted batch job 232542
The queue status will show your job as ACTIVE while its running, or WAITING if not.
SUMMARY OF JOBS FOR USER: <abattenh> ACTIVE JOBS-------------------- JOBID JOBNAME USERNAME STATE NODES REMAINING STARTTIME ================================================================================ 924965 simple abattenh Running 1 0:00:42 Sat Jun 3 21:33:31 WAITING JOBS------------------------ JOBID JOBNAME USERNAME STATE NODES WCLIMIT QUEUETIME ================================================================================ Total Jobs: 1 Active Jobs: 1 Idle Jobs: 0 Blocked Jobs: 0
If you don't see your simple job in either the ACTIVE or WAITING sections of your queue, it probably already finished – it should only run for a few seconds!
Notice in my queue status, where the STATE is Running, there is only one node assigned. Why is this, since there were 8 tasks?
Every job, no matter how few tasks requested, will be assigned at least one node. Each lonestar6 node has 128 physical cores, so each of the 8 tasks can be assigned to a different core.
Exercise: What files were created by your job?
filename wildcarding
You can look at one of the output log files like this:
cat cmd1.log
But here's a cute trick for viewing the contents all your output files at once, using the cat command and filename wildcarding.
cat cmd*.log
The cat command can take a list of one or more files. The asterisk ( * ) in cmd*.log is a multi-character wildcard that matches any filename starting with cmd then ending with .log.
You can also specify single-character matches inside brackets ( [ ] ) in either of the ways below, this time using the ls command so you can better see what is matching:
ls cmd[1234].log ls cmd[2-6].log
This technique is sometimes called filename globbing, and the pattern a glob. Don't ask me why – it's a Unix thing. Globbing – translating a glob pattern into a list of files – is one of the handy thing the bash shell does for you. (Read more about Pathname wildcards)
Exercise: How would you list all files starting with "simple"?
Here's what my cat output looks like. Notice the times are all nearly the same because all the tasks ran in parallel. That's the power of cluster computing!
Command 1 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:50 CDT 2023 Command 2 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:44 CDT 2023 Command 3 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:46 CDT 2023 Command 4 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:47 CDT 2023 Command 5 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:51 CDT 2023 Command 6 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:47 CDT 2023 Command 7 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:51 CDT 2023 Command 8 on c304-005.ls6.tacc.utexas.edu - Sat Jun 3 21:33:49 CDT 2023
echo
Lets take a closer look at a typical task in the simple.cmds file.
sleep 5; echo "Command 3 `date`" > cmd3.log 2>&1
The echo command is like a print statement in the bash shell. echo takes its arguments and writes them to standard output. While not always required, it is a good idea to put echo's output string in double quotes.
backtick evaluation
So what is this funny looking `date` bit doing? Well, date is just another Linux command (try just typing it in) that just displays the current date and time. Here we don't want the shell to put the string "date" in the output, we want it to execute the date command and put the result text into the output. The backquotes ( ` ` also called backticks) around the date command tell the shell we want that command executed and its standard output substituted into the string. (Read more about Quoting in the shell.)
# These are equivalent: date echo `date` # But different from this: echo date
output redirection
There's still more to learn from one of our simple tasks, something called output redirection.
Every command and Unix program has three "built-in" streams: standard input, standard output and standard error, each with a name, a number, and a redirection syntax.
Normally echo writes its string to standard output, but it could encounter an error and write an error message to standard error. We want both standard output and standard error for each task stored in a log file named for the command number.
sleep 5; echo "Command 3 `date`" > cmd3.log 2>&1
So in the above example the first '>' says to redirect the standard output of the echo command to the cmd3.log file. The '2>&1' part says to redirect standard error to the same place. Technically, it says to redirect standard error (built-in Linux stream 2) to the same place as standard output (built-in Linux stream 1); and since standard output is going to cmd3.log, any standard error will go there also. (Read more about Standard streams and redirection)
When the TACC batch system runs a job, all outputs generated by tasks in the batch job are directed to one output and error file per job. Here they have names like simple.e924965 and simple.o924965. simple.o924965 contains all standard output and simple.o924965 contains all standard error generated by your tasks that was not redirected elsewhere, as well as information relating to running your job and its tasks. For large jobs with complex tasks, it is not easy to troubleshoot execution problems using these files.
So a best practice is to separate the outputs of all our tasks into individual log files, one per task, as we do here. Why is this important? Suppose we run a job with 100 commands, each one a whole pipeline (alignment, for example). 88 finish fine but 12 do not. Just try figuring out which ones had the errors, and where the errors occurred, if all the standard output is in one intermingled file and all standard error in the other intermingled file!
Job parameters
Now that we've executed a really simple job, let's take a look at some important job submission parameters. These correspond to arguments to the launcher_creator.py script.
A bit of background. Historically, TACC was set up to cater to researchers writing their own C or Fortran codes highly optimized to exploit parallelism (the HPC crowd). Much of TACC's documentation is aimed at this audience, which makes it difficult to pick out the important parts for us.
The kind of jobs we biologists generally run are relatively new to TACC. They even have a special name for them: "parametric sweeps", by which they mean the same program running on different data sets.
In fact there is a special software module required to run our jobs, called the launcher module. You don't need to worry about activating the launcher module – that's done by the <job_name>.slurm script created by launcher_creator.py like this:
module load launcher
The launcher module knows how to interpret various job parameters in the <job_name>.slurm batch SLURM submission script and use them to create your job and assign its tasks to compute nodes. Our launcher_creator.py program is a simple Python script that lets you specify job parameters and writes out a valid <job_name>.slurm submission script.
launcher_creator.py
If you call launcher_creator.py with no arguments it gives you its usage description. Because it is a long help message, we may want to pipe the output to more, a pager that displays one screen of text at a time. Type the spacebar to advance to the next page, and Ctrl-c to exit from more.
# Use spacebar to page forward; Ctrl-c to exit launcher_creator.py | more
usage: launcher_creator.py [-h] -n NAME -t TIME_REQUEST [-j JOB_FILE]
[-b SHELL_COMMANDS] [-B SHELL_COMMANDS_FILE]
[-q QUEUE] [-a [ALLOCATION]] [-m MODULES]
[-M MODULES_FILE] [-w WAYNESS] [-N NUM_NODES]
[-e [EMAIL]] [-l LAUNCHER] [-s]
Create launchers for TACC clusters. Report problems to rt-
other@ccbb.utexas.edu
optional arguments:
-h, --help show this help message and exit
Required:
-n NAME, --name NAME The name of your job.
-t TIME_REQUEST, --time TIME_REQUEST
The time you want to give to your job. Format:
hh:mm:ss
Commands:
You must use at least one of these options to submit your commands for TACC.
-j JOB_FILE, --jobs JOB_FILE
The name of the job file containing your commands.
-b SHELL_COMMANDS, --bash SHELL_COMMANDS
A string of shell (Bash, zsh, etc) commands that are
executed before any parametric jobs are launched.
-B SHELL_COMMANDS_FILE, --bash_file SHELL_COMMANDS_FILE
A file containing shell (Bash, zsh, etc) commands that
are executed before any parametric jobs are launched.
Optional:
-q QUEUE, --queue QUEUE
The TACC allocation for job submission.
Default="development"
-a [ALLOCATION], -A [ALLOCATION], --allocation [ALLOCATION]
The TACC allocation for job submission. You can set a
default ALLOCATION environment variable.
-m MODULES, --modules MODULES
A list of module commands. The "launcher" module is
always automatically included. Example: -m "module
swap intel gcc; module load bedtools"
-M MODULES_FILE, --modules_file MODULES_FILE
A file containing module commands.
-w WAYNESS, --wayness WAYNESS
Wayness: the number of commands you want to give each
node. The default is the number of cores per node.
-N NUM_NODES, --num_nodes NUM_NODES
Number of nodes to request. You probably don't need
this option. Use wayness instead. You ONLY need it if
you want to run a job list that isn't defined at the
time you submit the launcher.
-e [EMAIL], --email [EMAIL]
Your email address if you want to receive an email
from Lonestar when your job starts and ends. Without
an argument, it will use a default EMAIL_ADDRESS
environment variable.
-l LAUNCHER, --launcher_name LAUNCHER
The name of the launcher script that will be created.
Default="<name>.slurm"
-s Echoes the launcher filename to stdout.
job name and commands file
Recall how the simple.slurm batch file was created:
launcher_creator.py -j simple.cmds -n simple -t 00:01:00 -a OTH21164 -q development
- The name of your commands file is given with the -j simple.cmds option.
- Your desired job name is given with the -n simple option.
- The <job_name> (here simple) is the job name you will see in your queue.
- By default a corresponding <job_name>.slurm batch file is created for you.
- It contains the name of the commands file that the batch system will execute.
queues and runtime
TACC resources are partitioned into queues: a named set of compute nodes with different characteristics. The main ones on ls6 are listed below. Generally you use development (-q development) when you are writing and testing your code, then normal once you're sure your commands will execute properly.
| queue name | maximum runtime | purpose |
|---|---|---|
| development | 2 hrs | development/testing and short jobs (typically has short queue wait times) |
| normal | 48 hrs | normal jobs (queue waits are often long) |
- In launcher_creator.py, the queue is specified by the -q argument.
- The default queue is development. Specify -q normal for normal queue jobs.
- The maximum runtime you are requesting for your job is specified by the -t argument.
- Format is hh:mm:ss
- Note that your job will be terminated without warning at the end of its time limit!
allocation and SUs
You may be a member of a number of different projects, hence have a choice which resource allocation to run your job under.
- You specify that allocation name with the -a argument of launcher_creator.py.
- If you have set an $ALLOCATION environment variable to an allocation name, that allocation will be used.
- When you run a batch job, your project allocation gets "charged" for the time your job runs, in the currency of SUs (System Units).
- SUs are related in some way to node hours, usually 1 SU = 1 node hour.
Jobs tasks should have similar expected runtimes
Jobs should consist of tasks that will run for approximately the same length of time. This is because the total node hours for your job is calculated as the run time for your longest running task (the one that finishes last).
For example, if you specify 100 commands and 99 finish in 2 seconds but one runs for 24 hours, you'll be charged for 100 x 24 node hours even though the total amount of work performed was only ~24 hours.
wayness (tasks per node)
One of the most confusing things in job submission is the parameter called wayness, which controls how many tasks are run on each compute node.
- Recall that there are 128 physical cores and 256 GB of memory on each compute node
- so theoretically you could run up to 128 commands on a node, each with ~2 GB available memory
- you usually run fewer tasks on a node, and when you do, each task gets more resources
| tasks per node (wayness) | cores available to each task | memory available to each task |
|---|---|---|
| 1 | 128 | ~256 GB |
| 2 | 64 | ~128 GB |
| 4 | 32 | ~64 GB |
| 8 | 16 | ~32 GB |
| 16 | 8 | ~16 GB |
| 32 | 4 | ~8 GB |
| 64 | 2 | ~4 GB |
| 128 | 1 | ~1 GB |
- In launcher_creator.py, wayness is specified by the -w argument.
- the default is 128 (one task per core)
- A special case is when you have only 1 command in your job.
- In that case, it doesn't matter what wayness you request.
- Your job will run on one compute node, and have all cores available.
Your choice of the wayness parameter will depend on the nature of the work you are performing: its computational intensity, its memory requirements and its ability to take advantage of multi-processing/multi-threading (e.g. bwa -t option or hisat2 -p option).
Bioinformatics programs generally perform substantial I/O, require more memory and fewer cores, so you'll generally want to run only a few tasks per node.
Wayness example
Let's use launcher_creator.py to explore wayness options. First copy over the wayness.cmds commands file:
# If $CORENGS is not defined: export CORENGS=/work/projects/BioITeam/projects/courses/Core_NGS_Tools cds mkdir -p core_ngs/slurm/wayness cd core_ngs/slurm/wayness cp $CORENGS/tacc/wayness.cmds .
Exercise: How many tasks are specified in the wayness.cmds file?
The wayness.cmds commands file consists of a number of identical lines that look like this:
sleep 3; echo "Command $LAUNCHER_JID of $LAUNCHER_NJOBS ($LAUNCHER_PPN per node) ran on node `hostname` core $LAUNCHER_TSK_ID" > cmd.$LAUNCHER_JID.log 2>&1
Create the batch submission script specifying a wayness of 4 (4 tasks per node), then submit the job and monitor the queue:
launcher_creator.py -j wayness.cmds -n wayness -w 4 -t 00:02:00 -a OTH21164 -q development sbatch wayness.slurm showq -u
Exercise: With 16 tasks requested and wayness of 4, how many nodes will this job require? How much memory will be available for each task?
Exercise: If you specified a wayness of 2, how many nodes would this job require? How much memory could each task use?
Look at the output file contents once the job is done.
cat cmd*log # or, for a listing ordered by node name (the 11th field) cat cmd*log | sort -k 11,11
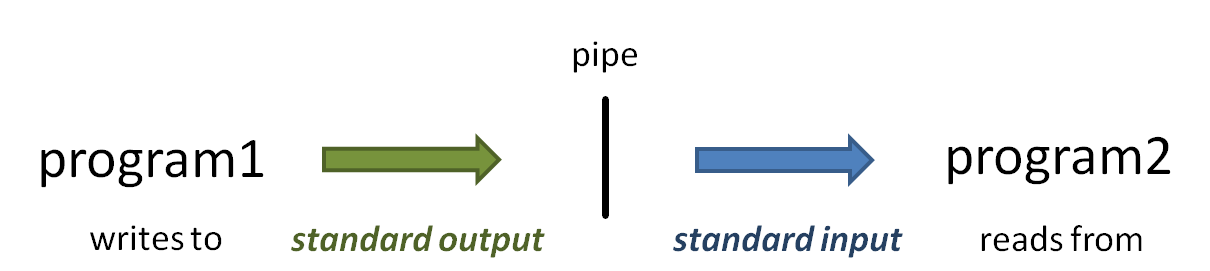
The vertical bar ( | ) above is the pipe operator, which connects one program's standard output to the next program's standard input.
(Read more about the sort command at Linux fundamentals: cut, sort, uniq, and more about Piping)
You should see something like output below.
Command 1 of 16 (4 per node) ran on node c303-005.ls6.tacc.utexas.edu core 0 Command 10 of 16 (4 per node) ran on node c304-005.ls6.tacc.utexas.edu core 9 Command 11 of 16 (4 per node) ran on node c304-005.ls6.tacc.utexas.edu core 10 Command 12 of 16 (4 per node) ran on node c304-005.ls6.tacc.utexas.edu core 11 Command 13 of 16 (4 per node) ran on node c304-006.ls6.tacc.utexas.edu core 12 Command 14 of 16 (4 per node) ran on node c304-006.ls6.tacc.utexas.edu core 13 Command 15 of 16 (4 per node) ran on node c304-006.ls6.tacc.utexas.edu core 14 Command 16 of 16 (4 per node) ran on node c304-006.ls6.tacc.utexas.edu core 15 Command 2 of 16 (4 per node) ran on node c303-005.ls6.tacc.utexas.edu core 1 Command 3 of 16 (4 per node) ran on node c303-005.ls6.tacc.utexas.edu core 2 Command 4 of 16 (4 per node) ran on node c303-005.ls6.tacc.utexas.edu core 3 Command 5 of 16 (4 per node) ran on node c303-006.ls6.tacc.utexas.edu core 4 Command 6 of 16 (4 per node) ran on node c303-006.ls6.tacc.utexas.edu core 5 Command 7 of 16 (4 per node) ran on node c303-006.ls6.tacc.utexas.edu core 6 Command 8 of 16 (4 per node) ran on node c303-006.ls6.tacc.utexas.edu core 7 Command 9 of 16 (4 per node) ran on node c304-005.ls6.tacc.utexas.edu core 8
Notice that there are 4 different host names. This expression:
cat cmd*log | awk '{print $11}' | sort | uniq -c
should produce output something like this (read more about piping commands to make a histogram)
4 c302-005.ls6.tacc.utexas.edu 4 c302-006.ls6.tacc.utexas.edu 4 c305-005.ls6.tacc.utexas.edu 4 c305-006.ls6.tacc.utexas.edu
Some best practices
Redirect task output and error streams
We've already touched on the need to redirect standard output and standard error for each task. Just remember that funny redirection syntax:
my_program input_file1 output_file1 > file1.log 2>&1
Combine serial workflows into scripts
Another really good way to work is to "bundle" a complex set of steps into a shell script that sets up its own environment, loads its own modules, then executes a series of program steps. You can then just call that script, probably with data-specific arguments, in your commands file. This multi-program script is sometimes termed a pipeline, although complex pipelines may involve several such scripts.
For example, you might have a script called align_bwa.sh (a bash script) or align_bowtie2.py (written in Python) that performs multiple steps needed during the alignment process:
- quality checking the input FASTQ file
- trimming or removing adapters from the sequences
- performing the alignment step(s) to create a BAM file
- sort the BAM file
- index the BAM file
- gather alignment statistics from the BAM file
The BioITeam maintains a set of such scripts in the /work/projects/BioITeam/common/script directory. Take a look at some of them after you feel more comfortable with initial NGS processing steps. They can be executed by anyone with a TACC account.
Use one directory per job
You may have noticed that all the files involved in our job were in one directory – the batch submissions file, commands file, log files our tasks wrote, and the launcher job output and error files. Of course you'll probably need input files too as well as output files .
Because a single job can create a lot of files, it is a good idea to use a different directory for each job or set of closely related jobs, maybe with a name similar to the job being performed. This will help you stay organized.
Here's an example directory structure
$SCRATCH/my_project
/original # contains or links to original fastq files
/fastq_prep # run fastq QC and trimming jobs here
/alignment # run alignment jobs here
/gene_counts # analyze gene overlap here
/test1 # play around with stuff here
/test2 # play around with other stuff here
Command files in each directory can refer to files in other directories using relative path syntax, e.g.:
cd $SCRATCH/my_project/fastq_prep ls ../original/my_raw_sequences.fastq.gz
Or create a symbolic link to the directory and refer to it as a sub-directory:
cd $SCRATCH/my_project/fastq_prep ln -s ../original fq ls ./fq/my_raw_sequences.fastq.gz
relative path syntax
As we have seen, there are several special "directory names" the bash shell understands:
- dot directory ( . ) refers to "here" or "the current directory"
- dot dot directory ( .. ) refers to "one directory up"
- tilde directory ( ~ ) refers to your Home directory
Try these relative path examples:
# navigate through the symbolic link in your Home directory cd ~scratch/core_ngs/slurm/simple ls ../wayness ls ../.. ls -l ~/.bashrc
(Read more about Absolute and relative pathname syntax)
Interactive sessions (idev)
So we've explored the TACC batch system. What if you want to do some interactive-style testing of your workflow?
Interactive sessions are available through the idev command as shown below. idev sessions are configured with similar parameters to batch jobs.
idev -m 60 -N 1 -A OTH21164 -p normal -r CoreNGS-Tue
Notes:
- -p normal requests nodes on the normal queue
- this is the default for our reservation, while the development queue is the normal default
- -m 60 asks for a 60 minute session
- -A OTH21164 specifies the TACC allocation/project to use
- -N 1 asks for 1 node
- --reservation=CoreNGS-Tue gives us priority access to TACC nodes for the class. You normally won't use this option.
When you ask for an idev session, you'll see output as shown below. Note that the process may repeat the "job status: PD" (pending) step while it waits for an available node.
-> Checking on the status of development queue. OK
-> Defaults file : ~/.idevrc
-> System : ls6
-> Queue : development (cmd line: -p )
-> Nodes : 1 (cmd line: -N )
-> Tasks per Node : 128 (Queue default )
-> Time (minutes) : 60 (cmd line: -m )
-> Project : OTH21164 (cmd line: -A )
-----------------------------------------------------------------
Welcome to the Lonestar6 Supercomputer
-----------------------------------------------------------------
--> Verifying valid submit host (login1)...OK
--> Verifying valid jobname...OK
--> Verifying valid ssh keys...OK
--> Verifying access to desired queue (development)...OK
--> Checking available allocation (OTH21164)...OK
Submitted batch job 235465
-> After your idev job begins to run, a command prompt will appear,
-> and you can begin your interactive development session.
-> We will report the job status every 4 seconds: (PD=pending, R=running).
-> job status: PD
-> job status: R
-> Job is now running on masternode= c302-005...OK
-> Sleeping for 7 seconds...OK
-> Checking to make sure your job has initialized an env for you....OK
-> Creating interactive terminal session (login) on master node c302-005.
-> ssh -Y -o "StrictHostKeyChecking no" c302-005
Once the idev session has started, it looks quite similar to a login node environment, except for these differences:
- the hostname command on a login node will return a login server name like login3.ls6.tacc.utexas.edu
- while in an idev session hostname returns a compute node name like c303-006.ls6.tacc.utexas.edu
- you cannot submit a batch job from inside an idev session, only from a login node
- your idev session will end when the requested time has expired
- or you can just type exit to return to a login node session