Objectives
In RNA-Seq, the abundance level of a gene is measured by the number of reads that map to that gene. Once the reads have been mapped to our reference, we must now count the number of reads that map to RNA units of interest to obtain gene/exon/transcript counts. Here, we shall look at different methods for doing this.
In our RNA-Seq analysis paths, here we will be exploring this path:
Count reads mapping to genes
Get set up
cds cd my_rnaseq_course cd day_1/tophat_results
Bedtools
Bedtools is a great utility for working with sequence features and mapped reads in BAM, BED, VCF, and GFF formats.
We are going to use it to count the number of reads that map to each gene in the genome. Load the module and check out the help for bedtools and the multicov specific command that we are going to use:
module load bedtools bedtools bedtools multicov
The multicov command takes a feature file (GFF) and counts how many reads are in certain regions from many input files. By default, it counts how many reads overlap the feature on either strand, but it can be made specific with the -s option. Unfortunately, this option only exists for the multicov command in a version of bedtools that is newer than the module on TACC, so we don't include it in the example command below.
Note: Remember that the chromosome names in your gff/gtf file should match the way the chromosomes are named in the reference fasta file used in the mapping step. For example, if reference file used for mapping contains chr1, chrX etc, the GFF/GTF file must also call the chromosomes as chr1, chrX and so on.
Let's double check this using grep.
grep '^>' reference/genome.fa
cut -f 1 reference/genes.gtf |sort|uniq -c
Let's fix the one chromosome that is differently named in the gtf file- mitochondrion.
sed 's/^dmel_mitochondrion_genome/M/' reference/genes.gtf > reference/genes.formatted.gtf cut -f 1 reference/genes.formatted.gtf |sort|uniq -c
In order to use the bedtools command on our data, do the following:
Warning: To submit to queue
nano commands.bedtools bedtools multicov -bams C1_R1_thout/accepted_hits.sorted.bam C1_R2_thout/accepted_hits.sorted.bam C1_R3_thout/accepted_hits.sorted.bam C2_R1_thout/accepted_hits.sorted.bam C2_R2_thout/accepted_hits.sorted.bam C2_R3_thout/accepted_hits.sorted.bam -bed reference/genes.formatted.gtf > gene_counts.gff #use ctrl+x to quit out of nano
launcher_creator.py -j commands.bedtools -n multicov -q normal -t 02:00:00 -a UT-2015-05-18 -l bedtools_launcher.slurm sbatch bedtools_launcher.slurm
HTseq
HTseq is another tool to count reads. bedtools has many many useful functions, and counting reads is just one of them. In contrast, HTseq is a specialized utility for counting reads. HTseq is very slow and you need to run multiple command lines in order to do the same job as what bedtools multicov did. However, if you are looking for more fine grained control over how to count genes, especially when a read overlaps more than one gene/feature, htseq-count would be an option.
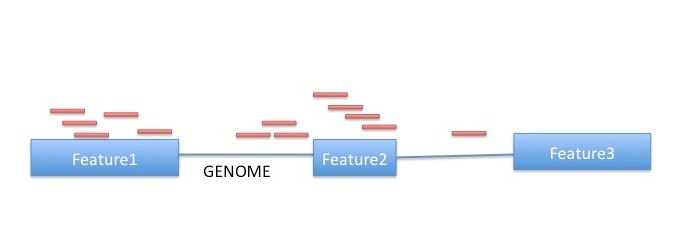
htseq-count has three modes for handling overlaps:
- mode union. This mode is recommended for most use cases.
- mode intersection-strict.
- mode intersection-nonempty.
Image from:http://www-huber.embl.de/users/anders/HTSeq/
module load htseq $TACC_HTSEQ_DIR/scripts/htseq-count
IMPORTANT NOTE: By default, htseq assumes your reads are stranded and will only count the reads that map in the same direction as the feature. If you have unstranded data or if you want to count reads mapping in all directions (maybe to detect antisense genes), use --stranded no option. If you have truseq data which uses dUTP method for creating stranded libraries, your reads will actually be in reverse direction when compared to the feature. So, use --stranded reverse.
$TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R1_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count1.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R2_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count2.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R3_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count3.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R1_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count4.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R2_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count5.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R3_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count6.gff join C1_count1.gff C1_count2.gff| join - C1_count3.gff | join - C2_count4.gff |join - C2_count5.gff|join - C2_count6.gff > gene_counts_HTseq.gff #if you have many samples, use for-loop and join
Warning: To submit to queue
nano commands.htseq $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R1_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count1.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R2_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count2.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C1_R3_thout/accepted_hits.sam reference/genes.formatted.gtf > C1_count3.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R1_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count4.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R2_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count5.gff $TACC_HTSEQ_DIR/scripts/htseq-count -m intersection-nonempty -i gene_id C2_R3_thout/accepted_hits.sam reference/genes.formatted.gtf > C2_count6.gff
launcher_creator.py -j commands.htseq -n htseq -q normal -t 02:00:00 -a UT-2015-05-18 -l htseq_launcher.slurm qsub htseq_launcher.slurm
AFTER THIS COMPLETES:
join C1_count1.gff C1_count2.gff| join - C1_count3.gff | join - C2_count4.gff |join - C2_count5.gff|join - C2_count6.gff > gene_counts_HTseq.gff #if you have many samples, use for-loop and join
Then take a peek at the results...
cds cd my_rnaseq_course/day_2/gene_expression_exercise
head gene_counts.gff wc -l gene_counts.gff
head gene_counts_HTseq.gff wc -l gene_counts_HTseq.gff
HTseq-count is strand-specific in default. Therefore, read counts for each gene in gene_counts_HTseq.gff are approximately a half counts in gene_counts.gff for the corresponding gene.
Other Gene Counting Options
If you want to perform all above operations in R enviornment, GRanges (along with Rsamtools) is a useful option. An example vignette is available here.
Let's look at how to check for differential expression now.